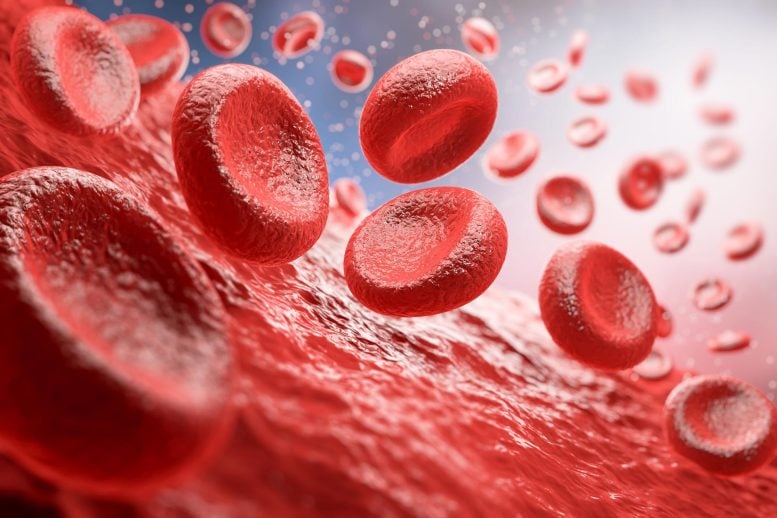
Scientists have uncovered a hidden layer of genetic regulation that helps explain why individuals with the same blood type can differ so dramatically at the molecular level.
A long-standing mystery in blood science has finally been solved, and the answer could make blood transfusions safer while shedding light on how our bodies fight disease.
Researchers at Lund University in Sweden have uncovered why people with the same blood type can carry vastly different amounts of key molecules on their red blood cells. The discovery, published in Nature Communications in 2023, explains a puzzle that has challenged scientists for decades and reveals hidden genetic controls that standard tests have missed.
Beyond Basic Blood Types
Blood groups are not just about whether you are A, B, AB, or O. They also depend on the number of specific molecules, called antigens, that sit on the surface of red blood cells. These molecules help the immune system distinguish between “self” and “foreign,” which is why precise matching is essential during transfusions.
Until now, scientists knew which genes produce these antigens, but not why their levels vary so widely between individuals with the same blood group.
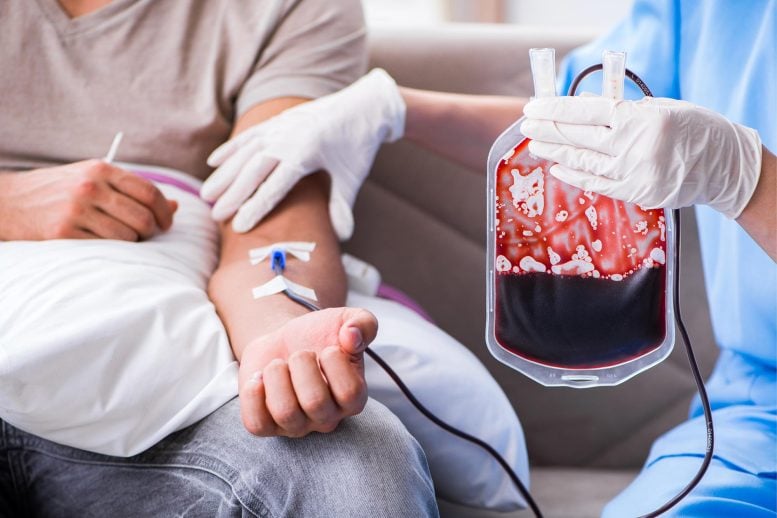
“This is important, because if you only have a couple of hundred blood group molecules per cell instead of a thousand or even a million molecules, then there is a risk that they maybe missed in a blood compatibility test, which can affect the safety of a blood transfusion,” explains Martin L Olsson, professor in Transfusion Medicine at Lund University and consultant within Clinical Immunology and Transfusion Medicine, Region Skåne, who led the study.
The Role of Genetic “Switches”
To solve the problem, the team looked beyond the genes themselves and focused on how those genes are controlled. They zeroed in on transcription factors, proteins that act like molecular switches by binding to specific DNA regions and regulating how strongly a gene is expressed.
Using a newly developed computational pipeline created by PhD student Gloria Wu, the researchers mapped nearly 200 of these binding sites across 33 blood group genes. This approach allowed them to predict where gene activity might be altered, something traditional genetic screening often overlooks.
They then tested their method on one of the most puzzling cases in transfusion medicine: the Helgeson blood group. Found in about 1% of the population, this rare variant is marked by unusually low levels of Complement Receptor 1 (CR1), a protein involved in immune defense. For years, its genetic cause remained unknown, and even DNA-based tests struggled to identify it.
“Margaret Helgeson was a medical technologist in Minneapolis in the 1970s who was trying to find compatible blood for a patient in need of a blood transfusion. Despite her best efforts, she could not find any suitable blood units. In desperation, she tested her own blood, and to her surprise, found it to be a match,” said Jill Storry, adjunct professor of experimental transfusion medicine at Lund University and a co-author of the study.
The new analysis revealed that the Helgeson variant is caused by a tiny change in a DNA sequence where a transcription factor is supposed to bind. Because the protein cannot attach properly, the CR1 gene is only weakly activated, leading to reduced levels of the molecule on red blood cells.
Evolutionary Trade-Offs and Disease Protection
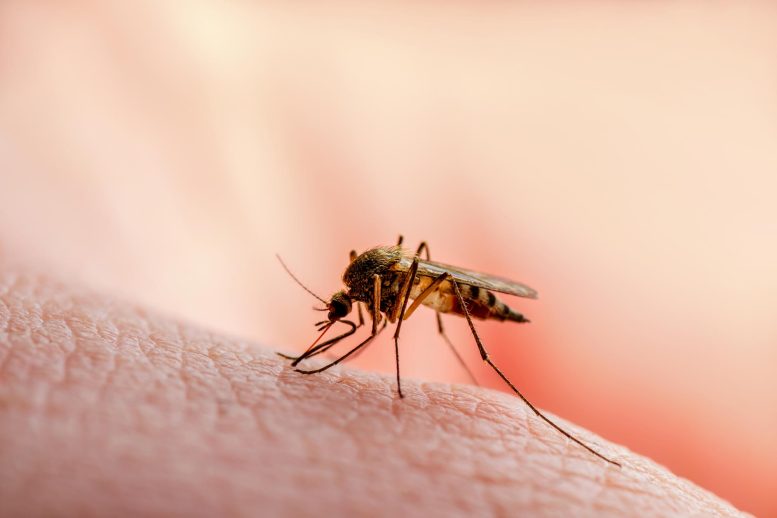
“Now the gene simply idles. In our study, we also showed this genetic variant to be more common in Thai blood donors compared with Swedish blood donors, which makes sense since we know from previous studies that a lower CR1 level is protective against malaria,” explains Martin L Olsson.
Lower CR1 levels appear to make it harder for malaria parasites to invade red blood cells, which may explain why the variant is more common in regions where the disease is widespread, such as Southeast Asia. In this way, a trait that complicates transfusion testing may also offer a survival advantage.
“Based on what we know now, we can improve the laboratory tests. Our goal is to update the existing DNA-based chip that is used for blood group tests with the new variant, which will result in a safer diagnostic test,” says Gloria Wu.
The team believes their data-driven approach can now be applied to many other blood groups and to diseases in which gene regulation plays a role. Follow-up studies already support that broader potential.
Expanding the Genetic Map
In 2024, one follow-up study in Transfusion Medicine and Hemotherapy showed that the same kind of hidden regulatory change can also affect the clinically crucial RhD blood group, identifying a novel mutation that disrupts a GATA1 binding site and reduces RhD expression to extremely low “Del” levels.
Another study in Transfusion expanded the researchers’ computational pipeline by combining transcription factor binding data with epigenetic markers and chromatin accessibility maps, revealing 814 potential regulatory sites across 47 blood group genes and confirming that combinations of factors such as GATA1 and KLF1 help control gene activity.
“Much of our research on blood groups now uses a combination of data-based predictive tools that can point us to the right experiment to test in the lab. The next challenge is to better understand the function of blood groups by connecting the information from large databases on how diseases affect people differently depending on their blood group,” concludes Martin L Olsson.
References:
“Elucidation of the low-expressing erythroid CR1 phenotype by bioinformatic mining of the GATA1-driven blood-group regulome” by Ping Chun Wu, Yan Quan Lee, Mattias Möller, Jill R. Storry and Martin L. Olsson, 17 August 2023, Nature Communications.
DOI: 10.1038/s41467-023-40708-w
“A Bioinformatically Initiated Approach to Evaluate GATA1 Regulatory Regions in Samples with Weak D, Del, or D– Phenotypes Despite Normal RHD Exons ” by Eunike C. McGowan, Ping Chun Wu, Åsa Hellberg, Genghis H. Lopez, Catherine A. Hyland and Martin L. Olsson, 10 May 2024, Transfusion Medicine and Hemotherapy.
DOI: 10.1159/000538469
“Epigenetic dissection of human blood group genes reveals regulatory elements and detailed characteristics of KEL and four other loci” by Ping Chun Wu, Eunike C. McGowan, Yan Quan Lee, Sudip Ghosh, Jenny Hansson and Martin L. Olsson, 21 April 2024, Transfusion.
DOI: 10.1111/trf.17840
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.
2 Comments
Hi this was so fascinating and I have a question. How does this information for the Hemophilia and other clotting disorders factor into your research? Could it help with blood compatibility when needing a factor 8 infusion?
Thanks for your time,
Laura Williamson
nice