Movies such as ‘X-Men,’ ‘Fantastic Four,’ and ‘The Guardians,’ which showcase vibrant mutant heroes, have captivated global audiences. Recently, a high-throughput genetic screening of meiotic crossover rate mutants in Arabidopsis thaliana garnered the interest of the academic community by unraveling a century-old mystery in the life sciences.
A research team, consisting of Professor Kyuha Choi, Dr. Jaeil Kim, and PhD candidate Heejin Kim from the Department of Life Sciences at Pohang University of Science and Technology (POSTECH), has achieved a remarkable feat by unveiling the molecular mechanism responsible for crossover interference during meiosis, a biological pattern at the chromosome level. The findings of this research were published on February 20 in Nature Plants, an international journal in the field of life sciences.
The Role of Meiosis in Genetic Diversity
In sexually reproducing organisms, individuals resemble their parents or siblings. Despite the striking similarities, it’s crucial to recognize that absolute identicalness is unattainable. This variation is attributed to the process of meiosis, which generates reproductive cells like sperm and eggs in animals or pollen and ovules in plants. Unlike somatic cell division, which duplicates and divides the genome identically, meiosis creates genetically diverse reproductive cells through a mechanism known as crossover.
Meiosis and crossover play pivotal roles in biodiversity and have significant implications in breeding where the selection and cultivation of superior traits in crops occur. Typically, most animal and plant species exhibit a minimum of one and a maximum of three crossovers per a pair of homologous chromosomes.
The ability to control the number of these crossovers could lead to cultivating crops with specific desired traits. However, achieving such control has been challenging due to the ‘phenomenon of crossover interference.’ Crossover interference, where one crossover inhibits the formation of another crossover nearby along the same chromosome, was initially identified by fruit fly geneticist Hermann J. Muller in 1916. Despite researchers’ persistent efforts over the past century since its discovery, it is only recently that the mechanisms underlying crossover interference have started to unveil their secrets.
Breakthrough in Understanding Crossover Interference
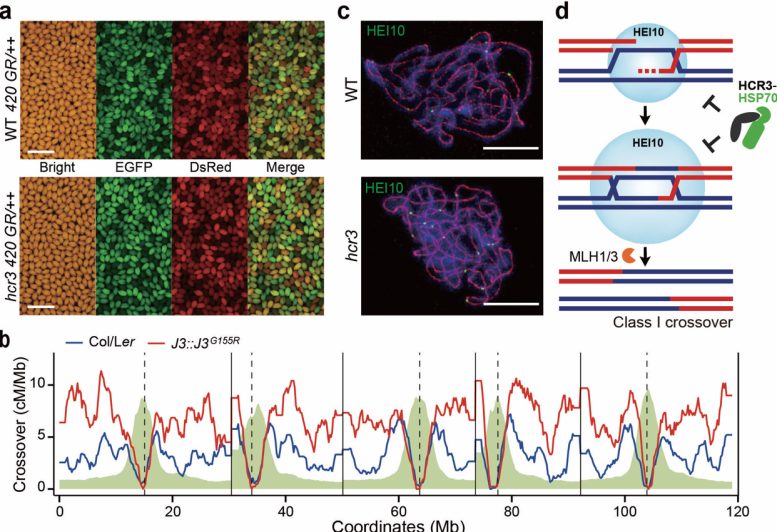
In this research, the team utilized a high-throughput fluorescent seed scoring method to directly measure crossover frequency in Arabidopsis plants. Through a genetic screen, they identified a mutant named hcr3 (high crossover rate3) that exhibited an increased crossover rate at the genomic level. Further analysis revealed that the elevated crossovers in hcr3 was attributed to a point mutation in the J3 gene, which encodes a co-chaperone related to HSP40 protein.
This research demonstrated that a network involving HCR3/J3/HSP40 co-chaperone and the chaperone HSP70 controls crossover interference and localization by facilitating the degradation of the pro-crossover protein, HEI10 ubiquitin E3 ligase. The application of genetic screen approaches to uncover the crossover interference and inhibition pathway successfully addressed a century-old puzzle in the life sciences.
POSTECH Professor Kyuha Choi stated, “Applying this research to agriculture will enable us to rapidly accumulate beneficial traits, thereby reducing breeding time.” He expressed optimism by saying, “We hope this research will contribute to the breeding of new varieties and identification of useful natural variations responsible for desirable traits such as disease and environmental stress resistance, improved productivity, and high-value production.”
Reference: “Control of meiotic crossover interference by a proteolytic chaperone network” by Heejin Kim, Jaeil Kim, Namil Son, Pallas Kuo, Chris Morgan, Aurélie Chambon, Dohwan Byun, Jihye Park, Youngkyung Lee, Yeong Mi Park, John A. Fozard, Julie Guérin, Aurélie Hurel, Christophe Lambing, Martin Howard, Ildoo Hwang, Raphael Mercier, Mathilde Grelon, Ian R. Henderson and Kyuha Choi, 20 February 2024, Nature Plants.
DOI: 10.1038/s41477-024-01633-y
The research was conducted with support from the Basic Research Program in Science and Engineering and the Mid-Career Researcher Program of the National Research Foundation of Korea, the Next-Generation BioGreen 21 Program of the Rural Development Administration, the Suh Kyungbae Foundation, and the Samsung Science & Technology Foundation.
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.
3 Comments
It’s not so much a question about cell division.
Thanks very for giving me the opportunity to learn new things in advance.
I need more understanding on miosis and mitosis