SSRgenotyper streamlines SSR genotyping by using sequencing data, boosting accuracy and eliminating lab-based methods.
SSRgenotyper is a newly developed, free bioinformatic tool that allows researchers to digitally genotype sequenced populations using simple sequence repeats (SSRs), a task that previously required time-consuming lab-based methods.
Reporting in a recent issue of Applications in Plant Sciences, the tool’s developers designed the program to seamlessly integrate with other applications currently used for the detection and analysis of SSRs.
The Genetic Power of Simple Sequence Repeats (SSRs)
Simple sequence repeats are short chains of repeating nucleotides that are prone to mutation. The variability of these DNA sequences makes them ideal for genetic analyses to distinguish between individuals and are often the marker of choice for paternity and forensic testing.
In research fields, SSRS have the added benefit of being selectively neutral, meaning they don’t code for any physical traits and therefore aren’t subject to most types of natural selection, making them an excellent tool to study populations without the obscuring effects of convergent evolution.
Recent advances in next-generation sequencing have helped streamline the process of SSR identification, especially in model organisms or groups with an available reference genome assembly. As technology continues to improve and sequencing costs decrease, sequencing large portions of a genome for the purposes of SSR analysis, even in non-model organisms, is becoming more feasible and widespread in the scientific literature.
From Gel to Genome: A Shift in Genotyping Methods
However, the process of genotyping — determining which individuals have which alleles — still relies predominantly on visualizing amplified DNA on an electrophoresis gel, an involved and potentially hazardous process, as DNA fragments are often stained with carcinogenic chemicals.
It also has the added issue of alleles being measured based on the size of the resulting bands, which is an estimate for the number of nucleotides in the amplified DNA fragment. Because there may be slight variations in the flanking regions that surround the SSRs of interest, and because there is no standardized method of determining an allele’s size using these methods, genotyping results from one experiment cannot be easily transferred or compared to those of another experiment.
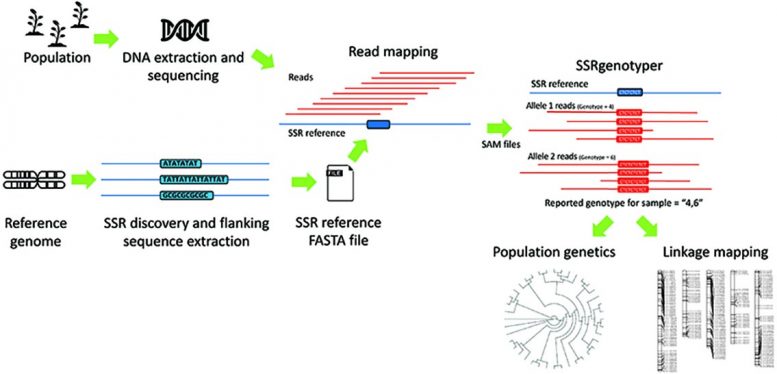
The development of SSRgenotyper renders such lab-based efforts obsolete. By working in tandem with other bioinformatic programs that detect SSRs in reference DNA and programs that align sequence data from target populations with the corresponding SSR reference file, SSRgenotyper is able to quickly genotype all SSRs for each individually sequenced sample.
“SSRgenotyper goes the next step by genotyping SSRs within sequenced populations — strictly from sequencing data (no PCR or electrophoresis),” said Jeff Maughan, a professor of Plant and Wildlife Sciences at Brigham Young University and senior author of the study. “The output from SSRgenotyper are files ready for population genetic analysis or linkage map formation.”
Streamlining Genetic Analysis: Benefits of Digital SSR Genotyping
Not only does the program reduce the amount of time and work required to genotype populations, it also solves the transferability problem inherent in electrophoresis estimates by directly counting the total number of base pairs in a given sequence repeat.
“Since the SSRs are genotyped based on the number of repeated motifs at the SSR locus and not on the PCR product size, the allele calls are standardized and transferable from project to project or from lab to lab,” said Maughan.
The program, which is coded in Python 3, requires only three positional arguments to run, provides the option to specify several conditional arguments (such as percentage thresholds for heterozygosity, the size of the flanking regions, and for the removal of spurious alleles), and can be performed on a regular desktop computer.
Once complete, SSRgenotyper generates multiple file types, including basic summary and statistical files, as well as a .pop, a .map, and an alignment file formatted for use in additional programs to facilitate downstream analyses.
Proven Performance with High Accuracy in Test Species
As a proof of concept, Maughan and his colleagues tested SSRgenotyper’s accuracy at correctly determining an individual’s genotype by running the program on publicly available sequences of quinoa (Chenopodium quinoa) and the oat species Avena atlantica. The resulting accuracy rate was 97% or greater, which increased with the inclusion of additional sequence reads.
With the continued development and efficiency of next-generation sequencing methods, tools like SSRgenotyper seem poised to reduce the amount of lab work required in genetic studies.
“Sequencing is already the method of choice in most genetic research projects,” said Maughan. “As costs continue to drop and new bioinformatic tools are developed, it is highly likely that future population genetics studies will be based solely on next-generation sequencing — completely avoiding the cumbersome tasks of PCR and electrophoresis.”
Reference: “SSRgenotyper: A simple sequence repeat genotyping application for whole‐genome resequencing and reduced representational sequencing projects” by Daniel H. Lewis, David E. Jarvis and Peter J. Maughan, 3 December 2020, Applications in Plant Sciences.
DOI: 10.1002/aps3.11402
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.