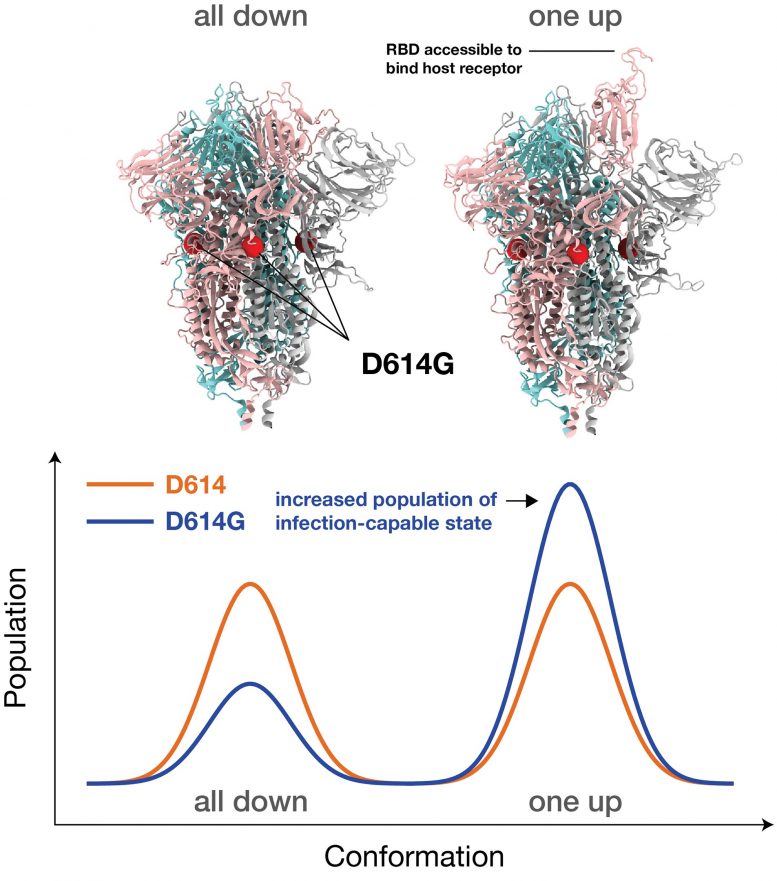
Dominant G-form Spike protein ‘puts its head up’ more frequently to latch on to receptors, but that makes it more vulnerable to neutralization.
Large-scale supercomputer simulations at the atomic level show that the dominant G form variant of the COVID-19-causing virus is more infectious partly because of its greater ability to readily bind to its target host receptor in the body, compared to other variantsThese research findings, led by a team from Los Alamos National Laboratory, shed light on the infection mechanism of the G form and its resistance to antibodies. This knowledge could contribute to future vaccine development.
Symmetrical Spike Protein Increases Binding Potential
“We found that the interactions among the basic building blocks of the Spike protein become more symmetrical in the G form, and that gives it more opportunities to bind to the receptors in the host — in us,” said Gnana Gnanakaran, corresponding author of the paper published recently in Science Advances. “But at the same time, that means antibodies can more easily neutralize it. In essence, the variant puts its head up to bind to the receptor, which gives antibodies the chance to attack it.”
Researchers knew that the variant, also known as D614G, was more infectious and could be neutralized by antibodies, but they didn’t know how. Simulating more than a million individual atoms and requiring about 24 million CPU hours of supercomputer time, the new work provides molecular-level detail about the behavior of this variant’s Spike.
Current vaccines for SARS-CoV-2, the virus that causes COVID-19, are based on the original D614 form of the virus. This new understanding of the G variant — the most extensive supercomputer simulations of the G form at the atomic level — could mean it offers a backbone for future vaccines.
The team discovered the D614G variant in early 2020, as the COVID-19 pandemic caused by the SARS-CoV-2 virus was ramping up. These findings were published in Cell. Scientists had observed a mutation in the Spike protein. (In all variants, it is the Spike protein that gives the virus its characteristic corona.) This D614G mutation, named for the amino acid at position 614 on the SARS-CoV-2 genome that underwent a substitution from aspartic acid, prevailed globally within a matter of weeks.
Molecular Dynamics of Spike Receptor Binding
The Spike proteins bind to a specific receptor found in many of our cells through the Spike’s receptor binding domain, ultimately leading to infection. That binding requires the receptor binding domain to transition structurally from a closed conformation, which cannot bind, to an open conformation, which can.
The simulations in this new research demonstrate that interactions among the building blocks of the Spike are more symmetrical in the new G-form variant than those in the original D-form strain. That symmetry leads to more viral Spikes in the open conformation, so it can more readily infect a person.
A team of postdoctoral fellows from Los Alamos — Rachael A. Mansbach (now assistant professor of Physics at Concordia University), Srirupa Chakraborty, and Kien Nguyen — led the study by running multiple microsecond-scale simulations of the two variants in both conformations of the receptor binding domain to illuminate how the Spike protein interacts with both the host receptor and with the neutralizing antibodies that can help protect the host from infection. The members of the research team also included Bette Korber of Los Alamos National Laboratory, and David C. Montefiori, of Duke Human Vaccine Institute.
The team thanks Paul Weber, head of Institutional Computing at Los Alamos, for providing access to the supercomputers at the Laboratory for this research.
Reference: “The SARS-CoV-2 Spike variant D614G favors an open conformational state” by Rachael A. Mansbach, Srirupa Chakraborty, Kien Nguyen, David C. Montefiori, Bette Korber, S. Gnanakaran, 16 April 2021, Science Advances.
DOI: 10.1126/sciadv.abf3671
Funding: The project was supported by Los Alamos Laboratory Directed Research and Development project 20200706ER, Director’s Postdoctoral fellowship, and the Center of Nonlinear Studies Postdoctoral Program at Los Alamos.
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.