For the first time, scientists from the Stowers Institute for Medical Research and collaborators have described the structure of an endogenously sourced, functioning neuronal amyloid at atomic resolution. The amyloid is composed of self-aggregated Orb2, the fruit fly version of the mRNA-binding cytoplasmic polyadenylation element-binding (CPEB) protein, which has been linked to long-term memory storage. The results of this work, published online on March 13, 2020, in Science, have some very interesting implications.
“We thought the usual way amyloids arise is when a protein, having for some reason gone bad or become misfolded, cues amyloid formation in a stochastic, unregulated process,” explains Kausik Si, Ph.D., Stowers investigator, and associate scientific director. “We found instead that amyloids can form at a very specific time, in a very specific cell, in a very specific way.”
Amyloids are usually understood in the context of their associated neurotoxic or degenerative states. In the case of Alzheimer’s, Parkinson’s, Huntington’s, and Creutzfeldt-Jakob diseases, proteins aggregate in an aberrant way to form stable, insoluble deposits that wreak havoc in the nervous system.
However, in 2003, Si and his mentor at the time, Eric Kandel, MD, discovered the existence of an amyloid with adaptive functions in the nervous system while studying CPEB in the sea slug Aplysia californica. Through subsequent studies in mice and the fruit fly Drosophila melanogaster, Si and others showed that in fact, the unique attributes of CPEB and Orb2 protein self-aggregates (amyloids), were essential to their proper functioning at synapses.
These studies have indicated that CPEB/Orb2 exists in different functional and structural states in the brain. The primary form is a monomer that represses translation at synapses, but as memory is formed, these monomers self-assemble into biochemically active aggregates that promote synaptic translation. This transition is necessary for memory persistence.
“They postulated that the amyloid state was likely important, but until this current work, it hadn’t been demonstrated definitively that the physical state of the protein is an amyloid,” explains Ruben Hervas, Ph.D., first author of the paper and a senior research associate in the Si Lab.
Hervas studied disease-causing amyloids during graduate school. “I wanted to apply my biophysical background to study the structural features of other amyloid-forming proteins — the good ones,” he says.
After Hervas joined the lab in 2016, he and Si agreed early on that the best way to study the structure of Orb2 was to use protein from an endogenous source (many fruit flies), rather than recombinant Orb2 extracted from bacteria — although it would be much more difficult.
Recombinant Orb2, unlike endogenous Orb2, is not biochemically active, explains Hervas. “The original environment, the nervous system, is important to preserve the active conformation of the protein, as well as its most interesting properties — relating to its ability to self-aggregate and form a biochemically active amyloid structure linked to memory.”
Hervas and the team faced an additional hurdle. “Because the protein is so large, about 700 amino acids, it is not easy to crystallize. For example, amyloid structures solved by X-ray crystallography thus far have only used peptides composed of few amino acids.” So, Hervas continues, “cryo-electron microscopy (EM) offered the opportunity to solve this kind of structure.”
In the current work, Hervas and colleagues purified Orb2 from approximately three million adult Drosophila heads. In these samples, Orb2 existed as monomers, oligomers, or self-aggregated filaments (amyloids). The researchers first confirmed that the Orb2 filaments were capable of seeding further filament formation, and that the purified Orb2 forms were biochemically active– that is, capable of repressing protein translation in the monomeric state, and capable of activating protein translation in the oligomeric and filament states.
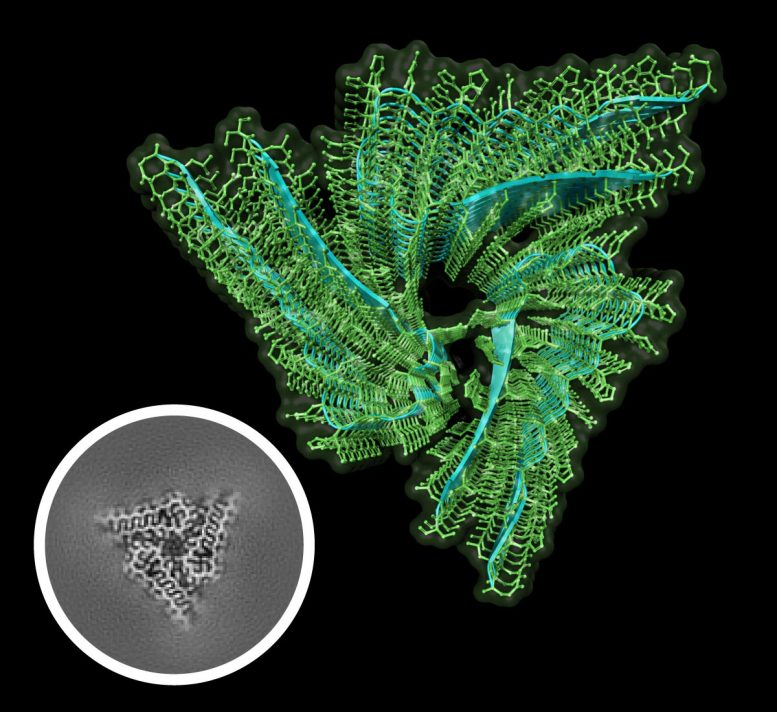
Their Herculean efforts paid off. Using cryo-EM and samples of over 97% purity, Hervas and colleagues were able to resolve the structure of endogenous Drosophila Orb2 at 2.6-angstrom resolution. They found that endogenous Orb2 formed threefold-symmetric amyloid filaments that were about 75 nanometers long. They confirmed that Orb2 aggregates in cross-beta amyloid structures and discovered that the hairpin-like fold of its protofilament cores, composed of 31 amino acids, pack via hydrophilic interfaces.
This work also provided a partial answer to a very important question — how is it possible that amyloids of a neuronal protein help store memory, when amyloids in general are associated with loss of memory? In their paper, the authors concluded that unlike the hydrophobic core of pathogenic amyloids, the hydrophilic core of Orb2 filaments suggests how some neuronal amyloids could be a stable yet regulatable substrate of memory.
“This finding changes the way we think about proteins folding and assembly,” says Si. “The way we think about biology now is: you have a gene, you make a string of amino acids, and once a polypeptide is made it is programmed to go into one conformation because there is a specific function attached to it. If it deviates from that path, a system is there to remove it. But this work suggests that sometimes a cell allows the same protein to form a completely different conformation. How does it do it? When does it do it?”
The conformational flexibility of some proteins opens the possibility that a protein may have more than one function depending on its conformational state — a phenomenon Si and his lab members will be keen to study moving forward.
While memory assays in Drosophila are informative, Hervas and colleagues are next looking forward to determining whether such functional amyloids also exist in the nervous system of mice and humans, especially because the nervous system of humans is particularly susceptible to amyloid-based diseases. There are four CPEB isoforms, each one with several variants, in mice and humans, explains Hervas. “Some specific isoforms in mice are also important for memory consolidation, and these are almost identical in protein sequence to corresponding human isoforms.”
“We are starting to work with CPEB proteins extracted from the human medial temporal lobe, a structure where long-lasting memories reside. We intend to solve the structure, and then potentially make the connection between memory and protein structure using mouse models.”
“The brain is one of the systems that evolved and diversified the most as organisms became more complex,” Si reflects. “But if you look at the number of proteins involved, that actually did not dramatically change. One possibility is that more complex biological systems use this conformational space to create more functions. Instead of creating new proteins, just being in a different shape could create new function.”
Other coauthors of this work include Younshim Park from the Stowers Institute; Michael J. Rau and James A.J. Fitzpatrick, Ph.D., from Washington University in St. Louis; and Wenjuan Zhang, Ph.D., Alexey G. Murzin, Ph.D., and Sjors H.W. Scheres, Ph.D., from the MRC Laboratory of Molecular Biology.
This work was funded in part by the Stowers Institute for Medical Research and by the UK Medical Research Council (award MC_UP_A025_1013, to S.H.W.S.).
Lay Summary of Findings
Proteins are generally thought to adopt a single shape, or “fold,” and the shape of a protein determines its function. However, some proteins can adopt a secondary shape, called “amyloid,” as they self-assemble. Historically, amyloids have been thought of as non-functional and even disease-causing, especially in the brain, as in the case of Alzheimer’s disease.
Surprisingly, though, work by Kausik Si, PhD, of the Stowers Institute for Medical Research, and others has indicated that a protein necessary for long-term memory formation and persistence, called Orb2, may be able to adopt the shape of an amyloid as part of its normal and necessary function. However, it had not been established that Orb2 exists in the brain as a bona fide amyloid, as its structure was unknown, until now.
A report from the Si Lab and collaborators, published online on March 13, 2020, in Science, has characterized, at the atomic-level resolution, the structure of Orb2 purified from its native source (the brains of millions of fruit flies) and demonstrated that it does indeed exist as a functional amyloid within the brain. This study, led by Ruben Hervas, Ph.D., is the first time an amyloid with a known biological function has ever been purified and described structurally from the brain. Interestingly, the amyloid (self-assembled) fold of Orb2 promotes the production of memory-related proteins, while the monomeric (single protein) fold of Orb2, inhibits the production of memory-related proteins, indicating that a protein can take on multiple functions by adopting multiple shapes.
These findings raise two interesting possibilities that will be further pursued in the Si Lab — the possibility that such a functioning neuronal amyloid also exists in humans, and the possibility that other proteins, known to have specific functions, are capable of achieving amyloid states to achieve alternate functions.
Reference: “Cryo-EM structure of a neuronal functional amyloid implicated in memory persistence in Drosophila” by Ruben Hervas, Michael J. Rau, Younshim Park, Wenjuan Zhang, Alexey G. Murzin, James A. J. Fitzpatrick, Sjors H. W. Scheres and Kausik Si, 13 March 2020, Science.
DOI: 10.1126/science.aba3526
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.