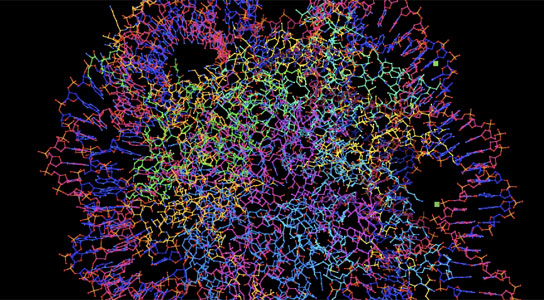
Protein folding
Proteins fold spontaneously into precise conformation, time after time, optimized by evolution. Yet given the number of contortions possible for chains of amino acids, dictating how a sequence will fold itself into a predictable structure has been a daunting task.
Researchers have been able to accomplish this feat. The scientists published their findings in the journal Nature. A team from David Baker’s laboratory at the University of Washington in Seattle has designed five proteins that fold reliably into predicted conformations. The synthesized proteins closely match the predicted structures.
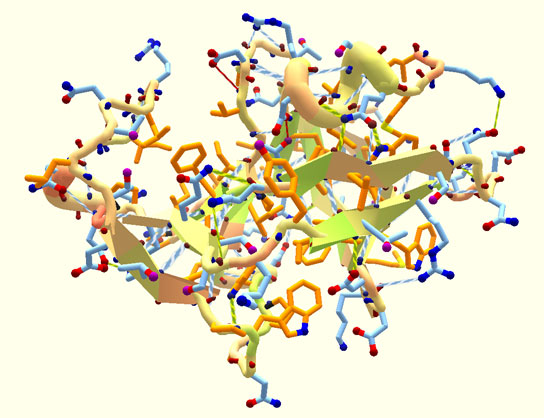
There’s only one previous example of a protein like this, and it was designed 10 years ago. Top7 was a one-off case, states Baker, a computational structural biologist.
They developed flexible sets of building blocks for nanoscale assembly, states Jeremy England, a molecular biophysicist at MIT. The work was led by the husband and wife team Nobuyasu and Rie Tatsumi Koga, protein engineers at the Baker group. After observing the backbone structures of thousands of proteins, they developed some intuitive rules they wanted to test.
Protein strands start from helices and secondary structures that fold into the final protein shape. These structures can be made to twist in one direction or another depending on the length of the loops that connect them. By choosing the right lengths, they could predict which way the proteins would fold.
The team developed a number of candidate sequences to fold into one of five structures. These structures were vetted by the group’s Rosetta@home program, which uses the home computers of volunteers to run protein-folding simulations. The sequences were folded hundreds of thousands of times. About 10% of the sequences had predicted structures that were stable enough. The winning sequences didn’t match any known naturally occurring proteins.
The proteins were synthesized and sent to Rutgers University in New Jersey to determine their structures using nuclear magnetic resonance (NMR) imaging.
The proteins are ideal because they are simple backbone constructs with every amino acid optimized to fold into the predicted, stable structure. That’s how they differ from natural proteins, whose folded structures represent a compromise between the requirements of optimum folding and biological function.
The reason why these proteins were designed from scratch is that natural proteins have been honed by evolution so precisely that it can be difficult to get the backbone to budge into another conformation to accommodate a new function.
Reference: “Principles for designing ideal protein structures” by Nobuyasu Koga, Rie Tatsumi-Koga, Gaohua Liu, Rong Xiao, Thomas B. Acton, Gaetano T. Montelione and David Baker, 7 November 2012, Nature.
DOI: 10.1038/nature11600






Doesn’t this create biohazards galore?
Creating completely new proteins of unknown biological properties seems quite dangerous to me.
The purpose of this experiment is to reveal the secrets of Protein folding which is engineered in the genetic code itself. The whole strand of DNA is neatly terminated by telomeres and folded into separate packs of chromosomes in the cell nucleus. The story of compartmentalization proceeds further into protein folding into desired shapes plane, ovoidal or other shapes in construction of tissues by a secret formula imbibed in the gene itself. When mRNA is brought inside the Ribosomes for making a Dye cast to read and construction starts with bits of tRNA threads, the protein formation is completed with the long thread churned out. This thread folds automatically into desired Protein shapes, which is imbibed in the code of gene itself. This experiment is attempting to work out the folding technique of protein and it is successful to some extent. It is purely a theoretical knowledge advancement. If artificial proteins are constructed, it may not be harmful but may supplement human nutrition one day. It is only the genetically modified products (BT products) that are to be concerned with.Thank You.