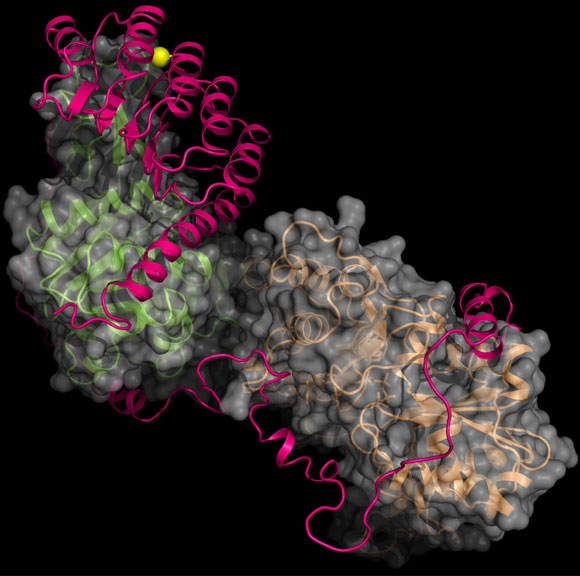
In a newly published study, a team of scientists from Rice University and Baylor College reveal the structural details of a protein seen as key to treating a neuromuscular disease.
Their success at obtaining a structural map of a protein known as leiomodin 2 (Lmod2) attached to two actin subunits offers a path forward for the study of nemaline myopathy, a hereditary disorder that weakens the muscles and can sometimes be fatal.
The work by biochemists Jianpeng Ma and Qinghua Wang and their colleagues appears this week in the Proceedings of the National Academy of Sciences. Ma holds a joint appointment at Rice and Baylor; Wang is at Baylor.
Proteins are large molecular chains of amino acids that perform many duties in the body. Knowing how the chains fold into functional proteins helps researchers determine how those involved in causing a disease might be addressed with drugs.
Obtaining actin’s structural details in atom-scale resolution had stumped many scientists, Ma said, because the protein’s rapid growth into an actin filament made it too long to be crystallized.
The Rice-Baylor effort came into focus 10 years ago when Ma realized that mutations can be designed to limit actin filament growth. Complexes with only two to four actin subunits would be short enough to be crystallized.
The researchers first solved actin’s long-standing resistance to analysis by making minor mutations that did not affect the actin subunits’ ability to form small molecular complexes called oligomers, but allowed it to be frozen in place and captured by X-ray crystallography, a century-old technique that reveals the position of every atom in a molecule. (Proteins are in constant motion as they carry out their functions. Researchers can study their three-dimensional forms only when they lock their atoms into position by crystallizing them.)
Ma and Wang were then most interested in the structure of Lmod2, a protein that nucleates actin and then mediates the formation of actin filaments within muscle cells. Bundles of filaments form fiber-like scaffolds that allow the elongated cells to expand and contract. Lmod2 is one of a family of proteins central to actin functions in many types of muscle cells, including cardiac and skeletal muscles.
“This protein is critical to biogenesis and length maintenance of actin filaments,” Wang said. “The filaments cannot be too short or too long.”
When Wang and Baylor postdoctoral researcher Xiaorui Chen mapped a mutation from another member of the leiomodin family known as Lmod3 observed in multiple patients onto Lmod2 in the lab, the resultant mutant lost its ability to nucleate actin filaments. That, Wang said, explains why the mutation can be lethal to patients.
“It has been a long and difficult process,” Chen said. “I am glad we persisted and succeeded.”
The work opens a window for researchers to treat nemaline myopathy by addressing the mutations, as well as a way to view mechanisms in other members of the leiomodin family. “This is just one example,” Wang said. “Because this protein has a large family, this structure not only helps us understand how this molecule works, it also helps understand how the family of this type of molecule works.”
In the process, Ma said, the team realized actin nucleators may also serve as an important component of signaling pathways in mammals that mediates the production of actin by acting as an “elegant” sensing mechanism. This possibility will be the subject of future study, he said.
Co-authors of the paper are Fengyun Ni, a postdoctoral associate at Baylor, and Elena Kondrashkina, a research scientist at the Synchrotron Research Center at Northwestern University. Ma is a professor of bioengineering and professor of biochemistry and cell biology at Rice and a professor of biochemistry at Baylor. Wang is an assistant professor of biochemistry at Baylor.
The National Institutes of Health and the Welch Foundation supported the research.
Reference: “Mechanisms of leiomodin 2-mediated regulation of actin filament in muscle cells” by Xiaorui Chen, Fengyun Ni, Elena Kondrashkina, Jianpeng Ma and Qinghua Wang, 28 September 2015, PNAS.
DOI: 10.1073/pnas.1512464112
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.
1 Comment
Am I missing something? Couldn’t the team find its structure just by using the AlphaFold computer program?