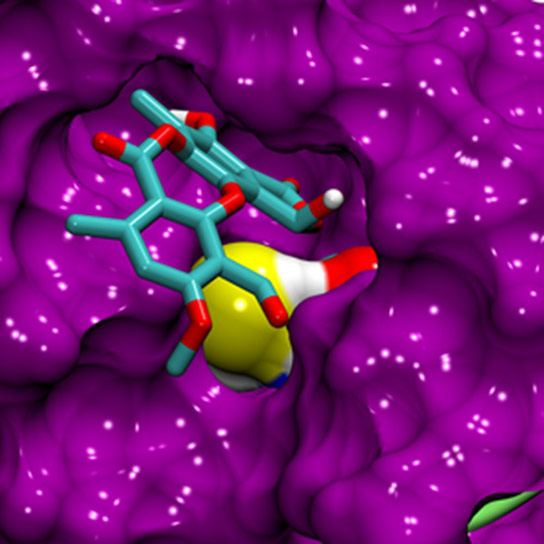
By using a computational method to capture the various shapes of the p53 protein, a protein that is implicated in nearly 40 percent of diagnosed cancer cases, scientists have identified a way to target the protein with cancer-fighting drugs.
University of California, Irvine biologists, chemists, and computer scientists have identified an elusive pocket on the surface of the p53 protein that can be targeted by cancer-fighting drugs. The finding heralds a new treatment approach, as mutant forms of this protein are implicated in nearly 40 percent of diagnosed cases of cancer, which kills more than half a million Americans each year.
In an open-source study published online this week in Nature Communications, the UC Irvine researchers describe how they employed a computational method to capture the various shapes of the p53 protein. In its regular form, p53 helps repair damaged DNA in cells or triggers cell death if the damage is too great; it has been called the “guardian of the genome.”
Mutant p53, however, does not function properly, allowing the cancer cells it normally would target to slip through control mechanisms and proliferate. For this reason, the protein is a key target of research on cancer therapeutics.
Within cells, p53 proteins undulate constantly, much like a seaweed bed in the ocean, making binding sites for potential drug compounds difficult to locate. But through a computational method called molecular dynamics, the UC Irvine team created a computer simulation of these physical movements and identified an elusive binding pocket that’s open only 5 percent of the time.
After using a computer to screen a library of 2,298 small molecules, the researchers selected the 45 most promising to undergo biological assays. Among these 45 compounds, they found one, called stictic acid, that fits into the protein pocket and triggers tumor-suppressing abilities in mutant p53s.
While stictic acid cannot be developed into a viable drug, noted study co-leader Peter Kaiser, professor of biological chemistry, the work suggests that a comprehensive screening of small molecules with similar traits may uncover a usable compound that binds to this specific p53 pocket.
“The discovery and pharmaceutical development of such a compound could have a profound impact on cancer treatments,” Kaiser said. “Instead of focusing on a specific form of the disease, oncologists could treat a wide spectrum of cancers, including those of the lung and breast.” He added that there is currently one group of experimental drugs – called Nutlins – that stop p53 degradation, but they don’t target protein mutations as would a drug binding to the newly discovered pocket.
The results are the culmination of years of labor by researchers with UC Irvine’s Institute for Genomics & Bioinformatics and the Chao Family Comprehensive Cancer Center.
“It’s been a large and complex multidisciplinary effort,” said Richard Lathrop, professor of computer science and co-leader of the study. “We’re working on the leading edge of what’s possible, and a variety of skills and expertise is required to make progress. Hopefully, our research eventually will lead to drugs that target many different forms of cancer.”
Hartmut Luecke, UC Irvine professor of molecular biology & biochemistry and physiology & biophysics, and Rommie Amaro, an assistant professor of computer science and pharmaceutical sciences who is now at UC San Diego, were other study co-leaders.
Additional UC Irvine team members included:
- Laboratory assistant Faezeh Salehi with the Department of Computer Science;
- Postdoctoral scholar Roberta Baronio, staff research associate Linda Hall, graduate student researcher Da-Wei Lin and undergraduate student Benjamin Chung with the Department of Biological Chemistry;
- Graduate student Brad Wallentine and postdoctoral scholar Chiung-Kuang Chen with the Department of Molecular Biology & Biochemistry;
- Richard Chamberlin, professor and chair of pharmaceutical sciences and professor of chemistry; and
- G. Wesley Hatfield, professor of microbiology & molecular genetics and chemical engineering & materials science.
Reference: “Computational identification of a transiently open L1/S3 pocket for reactivation of mutant p53” by Christopher D. Wassman, Roberta Baronio, Özlem Demir, Brad D. Wallentine, Chiung-Kuang Chen, Linda V. Hall, Faezeh Salehi, Da-Wei Lin, Benjamin P. Chung, G. Wesley Hatfield, A. Richard Chamberlin, Hartmut Luecke, Richard H. Lathrop, Peter Kaiser and Rommie E. Amaro, 29 January 2013, Nature Communications.
DOI: 10.1038/ncomms2361
Former UC Irvine postdoctoral researcher Christopher Wassman and Özlem Demir, a former UC Irvine postdoctoral scholar now at UC San Diego, also contributed.
This work was funded in part by the National Cancer Institute (grants R01CA112560 and T32CA009054), the National Institutes of Health (grants DP2-OD007237 and R01AI78000), the National Science Foundation (grants LRAC CHE060073N, CHE-0840513 and IIS-0326037) and the UC Irvine School of Medicine Dean’s Triumvirate Grant.
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.