A bacteria-only sugar has become the Achilles’ heel that could help defeat deadly superbugs.
Australian scientists have created a promising new approach for fighting deadly bacteria that no longer respond to antibiotics. The team designed antibodies that latch onto a sugar found only on bacterial cells, a discovery that could support a new class of immunotherapies aimed at multidrug-resistant infections acquired in hospitals.
The findings, published today (February 4) in Nature Chemical Biology, show that an antibody made in the laboratory was able to clear a normally fatal bacterial infection in mice. It works by locking onto a unique bacterial sugar and signaling the immune system to destroy the invading pathogen.
The research was co-led by Professor Richard Payne of the University of Sydney, working with Professor Ethan Goddard-Borger at WEHI and Associate Professor Nichollas Scott from the University of Melbourne and the Peter Doherty Institute for Infection and Immunity.
Professor Payne is also set to lead the newly announced Australian Research Council Centre of Excellence for Advanced Peptide and Protein Engineering. The center will build on discoveries like this one to speed progress toward real-world uses in biotechnology, agriculture, and conservation.
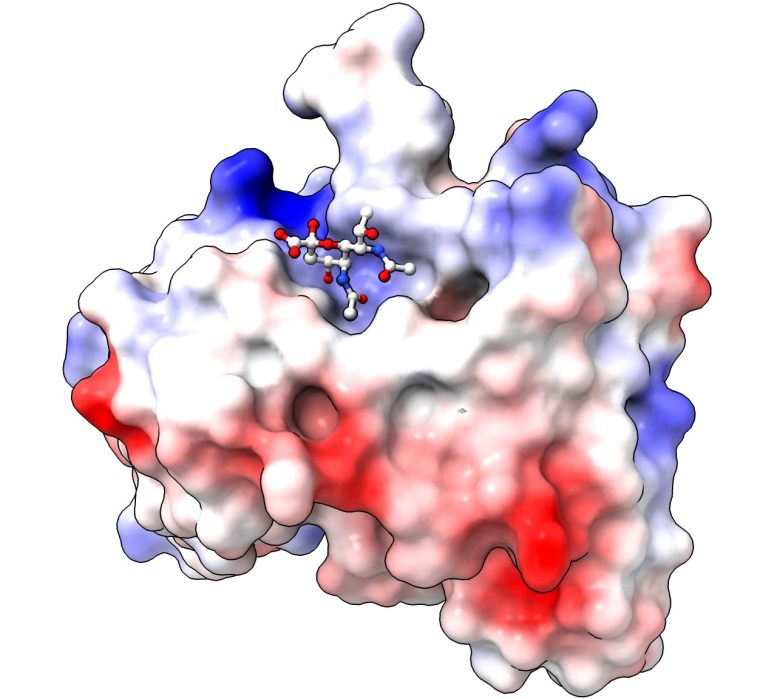
“This study shows what’s possible when we combine chemical synthesis with biochemistry, immunology, microbiology and infection biology,” Professor Payne said. “By precisely building these bacterial sugars in the lab with synthetic chemistry, we were able to understand their shape at the molecular level and develop antibodies that bind them with high specificity. That opens the door to new ways of treating some devastating drug-resistant bacterial infections.”
Why a Bacterial Sugar Makes an Ideal Target
The antibody targets a sugar molecule called pseudaminic acid. Although it looks similar to sugars found on human cells, this molecule is made only by bacteria. Many dangerous pathogens rely on it as a key part of their outer surface and use it to help avoid immune defenses.
Because the human body does not produce this sugar, it offers a highly specific and attractive target for developing immunotherapies that do not interfere with healthy cells.

Building Antibodies From the Ground Up
To take advantage of this vulnerability, the researchers first synthesized the bacterial sugar and sugar-decorated peptides entirely in the lab. This allowed them to map the molecule’s precise three-dimensional structure and see how it appears on bacterial surfaces.
Using that detailed understanding, the team created what they call a “pan-specific” antibody. It can recognize the same sugar across many different bacterial species and strains.
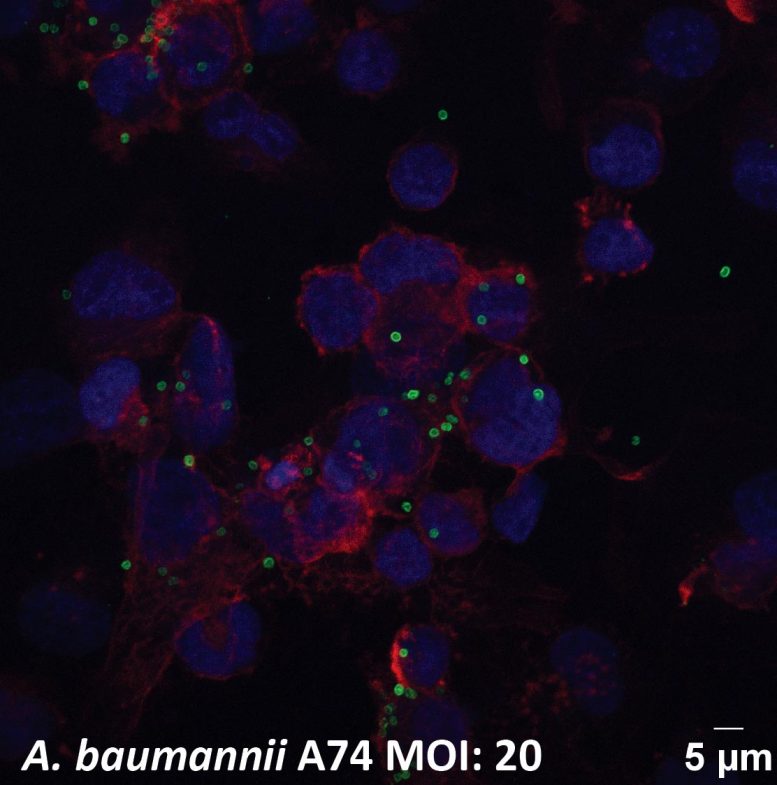
In mouse infection experiments, the antibody successfully eliminated multidrug-resistant Acinetobacter baumannii. This pathogen is a major cause of hospital-acquired pneumonia and bloodstream infections and is known for its ability to withstand multiple antibiotics.
“Multidrug-resistant Acinetobacter baumannii is a critical threat faced in modern healthcare facilities across the globe,” Professor Goddard-Borger said. “It is not uncommon for infections to resist even last-line antibiotics. Our work serves as a powerful proof-of-concept experiment that opens the door to the development of new life-saving passive immunotherapies.”

How Passive Immunotherapy Could Protect Patients
Passive immunotherapy involves giving patients ready-made antibodies so infections can be controlled quickly. This approach avoids the delay that comes with waiting for a person’s own immune system to mount a response.
The strategy can be used both to treat active infections and to prevent them. In hospital settings, it could help shield vulnerable patients in intensive care units from dangerous bacteria.
Associate Professor Scott noted that the antibodies also offer a new way to study how bacteria cause disease.
“These sugars are central to bacterial virulence, but they’ve been very hard to study,” he said. “Having antibodies that can selectively recognise them lets us map where they appear and how they change across different pathogens. That knowledge feeds directly into better diagnostics and therapies.”

Toward Clinical Use
Over the next five years, the researchers plan to turn their findings into antibody treatments ready for clinical testing against multidrug-resistant A. baumannii. Achieving this goal would remove one of the most dangerous members of the ESKAPE pathogens and mark an important step in the global effort to combat antimicrobial resistance.
“This is exactly the kind of breakthrough the new ARC Centre of Excellence is designed to enable,” Professor Payne said. “Our goal is to turn fundamental molecular insight into real-world solutions that protect the most vulnerable people in our healthcare system.”
Reference: “Uncovering bacterial pseudaminylation with pan-specific antibody tools” by Arthur H. Tang, Niccolay Madiedo Soler, Kristian I. Karlic, Leo Corcilius, Caitlin E. Clarke-Shepperson, Christopher Lehmann, Aleksandra W. Debowski, Ashleigh L. Dale, Lauren Zavan, Michelle Cielesh, Adedunmola P. Adewale, Karen D. Moulton, Lucy Li, Chenzheng Guan, Christopher McCrory, Maria Kaparakis-Liaskos, Benjamin P. Howden, Norelle L. Sherry, Ruohan Wei, Xuechen Li, Ruth M. Hall, Johanna J. Kenyon, Linda M. Wakim, Francesca L. Short, Danielle H. Dube, Stuart J. Cordwell, Mark Larance, Keith A. Stubbs, Glen P. Carter, Nichollas E. Scott, Ethan D. Goddard-Borger and Richard J. Payne, 4 February 2026, Nature Chemical Biology.
DOI: 10.1038/s41589-025-02114-9
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.