
Genomic ‘Tweezer’ Ushers In a New Era of Precision in Microbiome Research
Innovative method holds the potential to reshape our understanding of bacteria’s role in health and disease.
In a landmark study published today (January 4) in the journal Nature Methods, researchers at the Icahn School of Medicine at Mount Sinai have unveiled mEnrich-seq—an innovative method designed to substantially enhance the specificity and efficiency of research into microbiomes, the complex communities of microorganisms that inhabit the human body.
Unlocking the Microbial World with mEnrich-seq
Microbiomes play a crucial role in human health. An imbalance or a decrease in the variety of microbes in our bodies can lead to an increased risk of several diseases. However, in many microbiome applications, the focus is on studying specific types of bacteria in a sample, rather than looking at each type present. For example, when studying infectious diseases, researchers might only be interested in a few harmful gut bacteria, but they are mixed in with many other bacteria.
“Imagine you’re a scientist who needs to study one particular type of bacteria in a complex environment. It’s like trying to find a needle in a large haystack,” said Gang Fang, PhD, Professor of Genetics and Genomic Sciences and the study’s senior author. “mEnrich-seq essentially gives researchers a ‘smart tweezer’ to pick up the needle they’re interested in.”
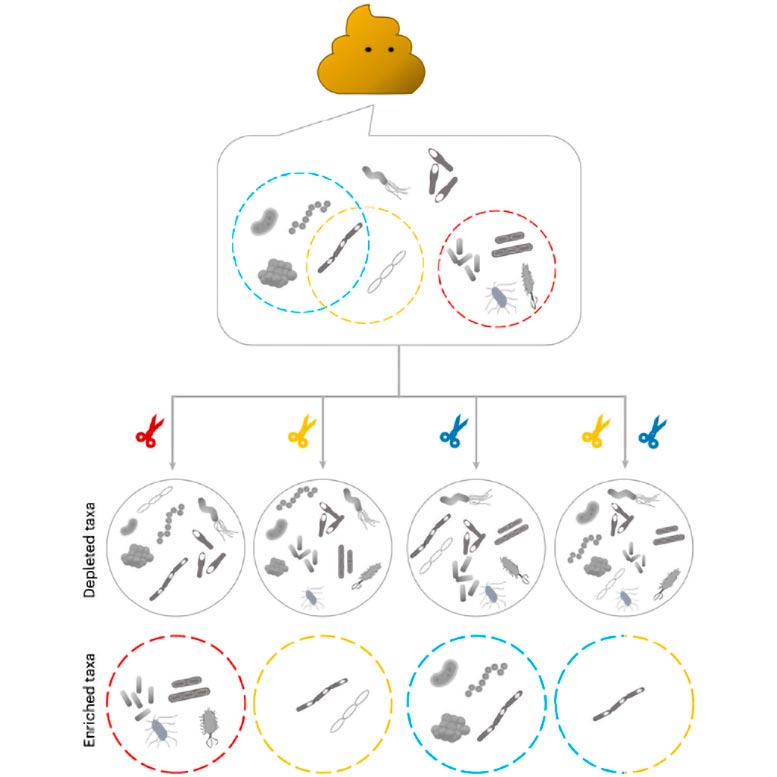
Once pulled out by the “smart tweezer,” researchers can assemble the genome(s) of the targeted bacteria, facilitating the study of diverse biomedical questions about them. This new strategy addresses a critical technology gap, as previously researchers would need to isolate specific bacterial strains from a given sample using culture media that selectively grow the specific bacterium—a time-consuming process that works for some bacteria, but not others. mEnrich-seq, in contrast, can directly recover the genome(s) of bacteria of interest from the microbiome sample without culturing.
mEnrich-seq effectively distinguishes bacteria of interest from the vast background by exploiting the “secret codes” written on bacterial DNA that bacteria use naturally to differentiate among each other as part of their native immune systems.
Transforming Research and Health Care
The advent of mEnrich-seq opens new horizons in various fields:
- Cost-Effectiveness: It offers a more economical approach to microbiome research, particularly beneficial in large-scale studies where resources may be limited.
- Broad Applicability: The method can focus on a wide range of bacteria, making it a versatile tool for both research and clinical applications.
- Medical Breakthroughs: By enabling more targeted research, mEnrich-seq could accelerate the development of new diagnostic tools and treatments.
“One of the most exciting aspects of mEnrich-seq is its potential to uncover previously missed details, like antibiotic resistance genes that traditional sequencing methods couldn’t detect due to a lack of sensitivity,” Dr. Fang added. “This could be a significant step forward in combating the global issue of antibiotic resistance.”
Indeed, as demonstrated as one of three applications in this study, the authors used mEnrich-seq to directly reconstruct pathogenic E. coli genomes from urine samples from patients with urinary tract infections, which allowed the comprehensive analysis of the antibiotic resistance genes in each genome.
In another application, the authors used mEnrich-seq to selectively construct the genomes of Akkermansia muciniphila, a bacterium that has been shown to have benefits in obesity and diabetes, among several other diseases, as well as a response to cancer immunotherapy. This bacterium is hard to culture, so mEnrich-seq can be a useful tool to reconstruct its genome in a culture-independent, sensitive, and cost-effective way, which may facilitate larger-scale association studies with different human diseases.
The Future of mEnrich-seq
Looking ahead, the team has ambitious plans for mEnrich-seq. They aim to refine the method to improve its efficiency further and to expand its range of applications. Collaborations with clinicians and healthcare professionals are also in the pipeline to validate the method’s utility in real-world settings.
“We envision mEnrich-seq as a sensitive and versatile tool in the future of microbiome studies and clinical applications,” said Dr. Fang.
Reference: “mEnrich-seq: methylation-guided enrichment sequencing of bacterial taxa of interest from microbiome” by Lei Cao, Yimeng Kong, Yu Fan, Mi Ni, Alan Tourancheau, Magdalena Ksiezarek, Edward A. Mead, Tonny Koo, Melissa Gitman, Xue-Song Zhang and Gang Fang, 4 January 2023, Nature Methods.
DOI: 10.1038/s41592-023-02125-1
This work was supported by a grant number R35 GM139655 from the National Institutes of Health.
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.