
What if one of the most iconic building blocks of life was misunderstood all along? A new study challenges long-standing assumptions about ancient proteins and their role in the origin of life.
How did life originate? Ancient proteins may hold important clues. Every living organism on Earth is built from proteins. Today, even the simplest life forms like single-celled organisms contain complex and highly structured proteins. But it wasn’t always this way.
For a long time, scientists believed that the earliest proteins began with a simple structural feature known as a “motif.”
This motif was thought to be the seed from which more complex proteins evolved. However, new research is challenging that idea. A global team of scientists, led by Lynn Kamerlin from Georgia Tech and Liam Longo from the Earth-Life Science Institute in Tokyo, has found that the motif might not be as central as once believed, at least not on its own.
“It’s probably an eroded molecular fossil, with its true nature having been overwritten over billions of years of evolution,” said Kamerlin. “This work completely reshapes how we think about proteins. It’s like trying to play protein Jeopardy! — now we need to rethink what the original question was.”
Prehistoric Proteins
It’s not hard to understand why this hypothesis was wrong for so long. The motif is associated with the element phosphorus, one of the key elements of life. Many of the earliest proteins bound to phosphorus-containing compounds. While these early proteins have different structures, they frequently share the same motif.
“For years, researchers took this to mean that today’s complex proteins came from the motif itself — that this tiny protein gave rise to entire families,” Longo said.
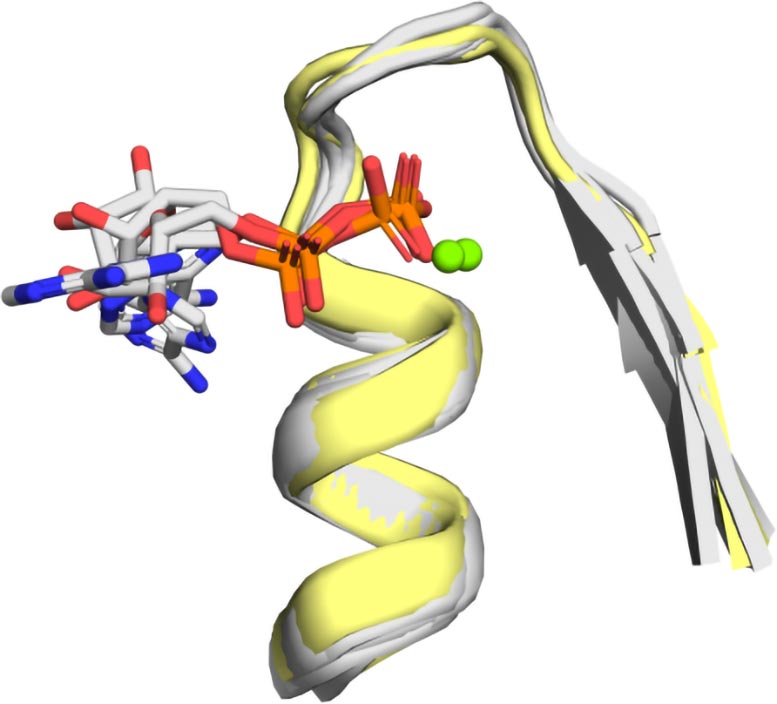
To discover the protein’s origins, the researchers pored over available data on protein crystal structures. Then they identified and characterized relevant proteins computationally. Although they recognized some of the protein’s similar structure in their modeling, the motif was not identical. They found that many different types of phosphate-binding proteins were possible. The idea that this motif was somehow special on its own was false.
“We don’t hypothesize that eyes gave rise to heads, even though nearly all heads have eyes; that’s because seeing involves interlocking systems,” Kamerlin said. “Our early peptide presents a similar instance. Only embedding within the larger system allows it to shine.”
Protein Possibilities
The researchers tested this work in water and methanol environments. Methanol mimics environments on Earth that may have less water around. The researchers found comparable protein motifs in this methanol environment, proving that the famous motif was not unique, but rather one of many possible motifs with similar properties. What was assumed to be a building block of early life is probably just a fossil fragment — and not the complete picture.
Kamerlin and Longo’s work helps their field determine not just how life started but also bolsters biotechnology advancements. A better understanding of how natural proteins evolved will help other researchers create artificial proteins, for everything from drug delivery to new vaccines.
The work is far from finished. Now that the researchers know this protein motif is one of many possible options, the question becomes: When did this motif become dominant, and what else could life have looked like? These questions will help the scientific world make discoveries that could benefit everyone.
Reference: “Redefining the Limits of Functional Continuity in the Early Evolution of P-Loop NTPases” by Andrey O Demkiv, Saacnicteh Toledo-Patiño, Encarnación Medina-Carmona, Andrej Berg, Gaspar P Pinto, Antonietta Parracino, Jose M Sanchez-Ruiz, Alvan C Hengge, Paola Laurino, Liam M Longo and Shina Caroline Lynn Kamerlin, 12 March 2025, Molecular Biology and Evolution.
DOI: 10.1093/molbev/msaf055
Funding from the Knut and Alice Wallenberg Foundation; the Okinawa Institute of Science and Technology Graduate University (OIST) with subsidy funding from the Cabinet Office, Government of Japan; and the National Academic Infrastructure for Supercomputing in Sweden.
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.
1 Comment
They essentially confirm that the P-loop motif evolved from a polyphosphate transporter and not the later phosphoryl transfer enzyme.
This aligns with what we know from the genetic code evolution, where transport properties were also central for amino acid recruitment. [S. Wehbi,A. Wheeler,B. Morel,N. Manepalli,B.Q. Minh,D.S. Lauretta,& J. Masel, Order of amino acid recruitment into the genetic code resolved by last universal common ancestor’s protein domains, Proc. Natl. Acad. Sci. U.S.A. 121 (52)]