
Researchers used molecular signatures in mobile DNA to trace the evolutionary origins of the cultivated strawberry genome.
Polyploid genomes form when entire genomes duplicate and combine through repeated hybridization events. This process has played a major role in the evolution of many crop plants.
However, understanding the internal structure of these genomes can be difficult, especially when the ancestral species that contributed to them are unknown. In a new study, researchers introduced a genome-wide method designed to unravel the complexity of polyploid genomes by analyzing evolutionary signals left behind by long terminal repeat retrotransposons.
The method compares similarity patterns of these genetic elements across chromosomes. This analysis allows scientists to reconstruct the organization of subgenomes and estimate when major genome merging events occurred.
When applied to cultivated octoploid strawberry, the approach revealed a stepwise evolutionary history shaped by multiple allopolyploidization events. The findings provide new insight into how complex plant genomes gradually assemble and diversify over millions of years.
Whole genome duplication has repeatedly reshaped plant genomes and contributed to evolutionary innovation, environmental adaptation, and crop diversity. In allopolyploid species, chromosomes originate from different ancestral genomes. These genomes combine into several subgenomes that continue to evolve and interact over time.
Identifying these subgenomes is critical for understanding how plant genomes develop and change. Traditional methods typically rely on known diploid ancestor species for comparison. However, many of these ancestors are either extinct or have not yet been identified.
Transposable elements, particularly long terminal repeat retrotransposons, accumulate in patterns that are unique to specific evolutionary lineages. These elements preserve molecular evidence of past genomic events. Despite this potential, researchers still lack widely applicable tools that can reliably convert these patterns into clear subgenome assignments. As a result, new approaches are needed to reconstruct polyploid genome evolution when ancestral genomes are unavailable.
A New Bioinformatic Framework for Polyploid Genomes
Scientists from the U.S. Department of Agriculture and several collaborating institutions have introduced a new computational framework that helps reconstruct the evolutionary history of complex polyploid genomes. The study was published in Horticulture Research.
The research team used a method called the serial similarity matrix, which analyzes long terminal repeat retrotransposons across the genome. Using this approach, they reexamined the genome of cultivated octoploid strawberry (Fragaria × ananassa). Their findings clarify the arrangement of strawberry subgenomes and reveal several ancient genome merging events that shaped the species. The results also address long standing questions about the evolutionary origin of the modern strawberry.
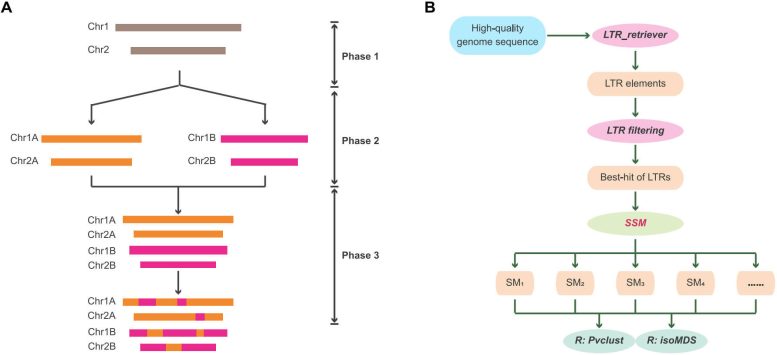
The researchers structured their approach around three major evolutionary phases: the period before ancestral species split from one another, the time during which those species evolved separately, and the stage after their genomes combined. Retrotransposons that expanded during the divergence phase carry signatures unique to each emerging lineage.
To capture these signals, the team calculated similarity matrices for retrotransposon sequences across chromosomes and examined how they clustered under different similarity thresholds. This stepwise analysis produced a “serial similarity matrix” that preserves evolutionary signals from different time periods within the genome.
Testing the Method in Known Polyploid Crops
Before turning to strawberry, the scientists validated their approach in well-studied allopolyploid crops such as teff and cotton. In these species, the method successfully distinguished known subgenomes and separated events that occurred before polyploidization from those that happened afterward.
They also applied the framework to artificially constructed polyploid genomes. These tests showed that the method responds predictably to differences in divergence time and to the abundance of transposable elements, supporting its reliability.
When applied to octoploid strawberry, the analysis identified four distinct subgenomes and uncovered three successive allopolyploidization events. These events likely occurred between approximately 3.1–4.2, 1.9–3.1, and 0.8–1.9 million years ago.
The findings indicate close evolutionary connections between two of the strawberry subgenomes and the diploid species Fragaria vesca and Fragaria iinumae. At the same time, the results challenge earlier models that suggested additional diploid progenitors. Instead, the study suggests that extinct or currently unsampled relatives probably played a role in shaping the strawberry genome, underscoring the intricate nature of polyploid evolution.
Transposable Elements as Evolutionary Time Stamps
“This work demonstrates how transposable elements can function as evolutionary time stamps embedded in plant genomes,” said one of the study’s senior authors. “By focusing on when and where these elements expanded, we can reconstruct genome history even when direct ancestral references are missing. This method provides a powerful new lens for studying polyploid crops and moves beyond reliance on incomplete progenitor data, offering a more objective and reproducible framework for evolutionary genomics.”
The implications extend well beyond strawberries. Many important crops, including wheat, cotton, and sugarcane, are polyploids with complicated evolutionary backgrounds. Resolving their subgenomes more accurately can strengthen gene annotation, improve trait mapping, and enhance comparative genomic studies.
By making it possible to reconstruct genome history without known ancestors, the serial similarity matrix approach broadens the set of tools available to scientists studying biodiversity, speciation, and adaptation. It also offers a practical framework for connecting evolutionary research with crop improvement and modern plant breeding.
Reference: “Deciphering octoploid strawberry evolution with serial LTR similarity matrices for subgenome partition” by Haomin Lyu, Shujun Ou, Won Cheol Yim and Qingyi Yu, 21 May 2025, Horticulture Research.
DOI: 10.1093/hr/uhaf132
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.
1 Comment
thanks