A bacterium preserved for millennia in cave ice has revealed unexpected resistance to modern antibiotics.
Bacteria are known for their ability to survive in some of the harshest places on Earth, from extreme heat to deep freezes. Ice caves are one such habitat, sheltering diverse microbial communities that scientists have only begun to explore for their genetic potential.
In Romania, researchers recently analyzed the antibiotic resistance of a bacterial strain that had been trapped for 5,000 years in the ice of an underground cave. Their findings, published in Frontiers in Microbiology, suggest that studying such ancient microbes could help scientists better understand how antibiotic resistance develops naturally and potentially guide new strategies to slow its spread.
“The Psychrobacter SC65A.3 bacterial strain isolated from Scarisoara Ice Cave, despite its ancient origin, shows resistance to multiple modern antibiotics and carries over 100 resistance-related genes,” said author Dr. Cristina Purcarea, a senior scientist at the Institute of Biology Bucharest of the Romanian Academy. “But it can also inhibit the growth of several major antibiotic-resistant ‘superbugs’ and showed important enzymatic activities with important biotechnological potential.”

Ancient resistance to modern medication
Psychrobacter SC65A.3 belongs to the genus Psychrobacter, a group of bacteria adapted to cold climates. While some members of this genus can infect humans or animals, they are also considered promising for biotechnology. However, little is known about how these bacteria respond to antibiotics.
“Studying microbes such as Psychrobacter SC65A.3 retrieved from millennia-old cave ice deposits reveals how antibiotic resistance evolved naturally in the environment, long before modern antibiotics were ever used,” said Purcarea.
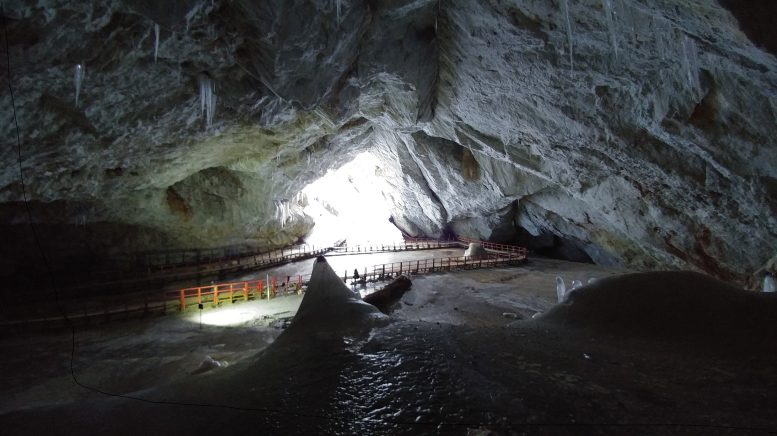
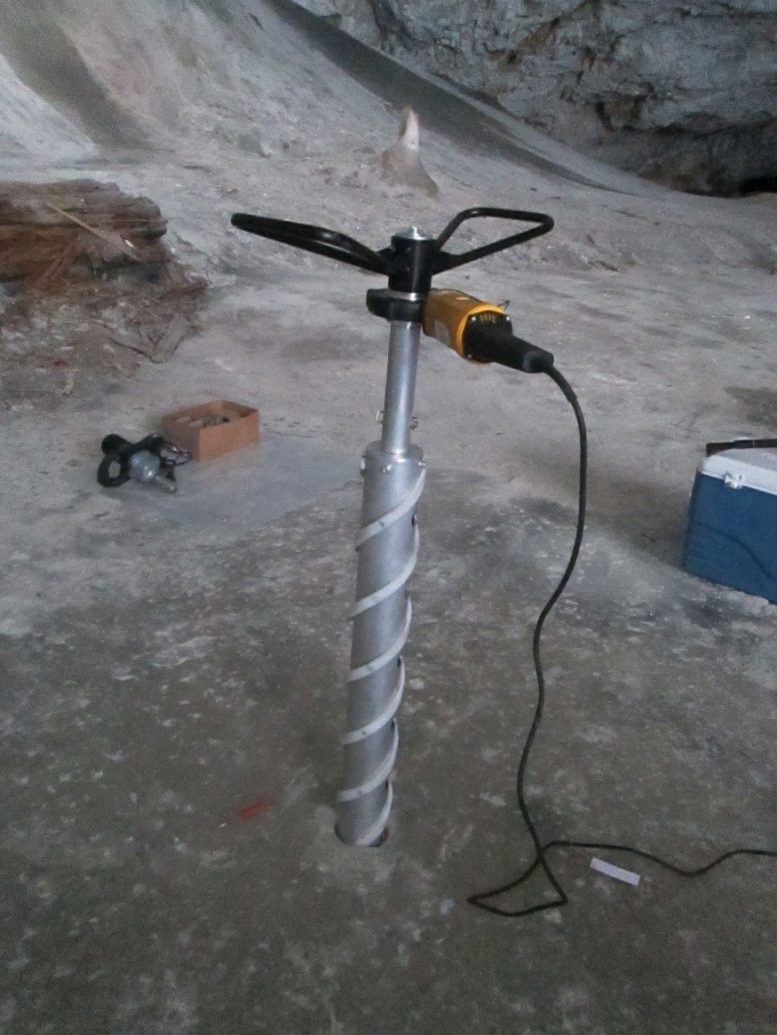
To investigate, the scientists drilled a 25-meter ice core from a section of the cave known as the Great Hall, capturing layers that span 13,000 years. The ice samples were sealed in sterile bags and kept frozen during transport to prevent contamination. In the laboratory, the team isolated bacterial strains and sequenced their genome to identify genes linked to cold survival, antimicrobial resistance, and antimicrobial activity.
The researchers then exposed the SC65A strain to 28 antibiotics from 10 different classes, including drugs commonly prescribed or reserved for serious infections. Some of these antibiotics are already known to be associated with resistance genes or mutations, allowing the team to compare genetic predictions with actual laboratory results.
“The 10 antibiotics we found resistance to are widely used in oral and injectable therapies used to treat a range of serious bacterial infections in clinical practice,” Purcarea pointed out. Diseases such as tuberculosis, colitis, and UTIs can be treated with some of the antibiotics that the researchers found resistance to, including rifampicin, vancomycin, and ciprofloxacin.

SC65A.3 is the first Psychrobacter strain shown to resist certain antibiotics, including trimethoprim, clindamycin, and metronidazole. These drugs are commonly used to treat UTIs as well as infections affecting the lungs, skin, bloodstream, and reproductive system. The strain’s resistance profile indicates that cold adapted bacteria may serve as reservoirs of resistance genes, which are segments of DNA that enable microbes to survive exposure to antibiotic treatments.
Risky potential
Bacterial strains like the one examined here hold both a threat and a promise. “If melting ice releases these microbes, these genes could spread to modern bacteria, adding to the global challenge of antibiotic resistance,” Purcarea said. “On the other hand, they produce unique enzymes and antimicrobial compounds that could inspire new antibiotics, industrial enzymes, and other biotechnological innovations.”
In the Psychrobacter SC65A.3 genome, the researchers found almost 600 genes with unknown functions, suggesting a yet untapped source for discovering novel biological mechanisms. Analysis of the genome also revealed 11 genes that are potentially able to kill or stop the growth of other bacteria, fungi, and viruses.
Such potential is becoming ever more important in a world where antibiotic resistance is a growing concern. Going back to ancient genomes and uncovering their potential highlights the important role the natural environment played in the spread and evolution of antibiotic resistance. “These ancient bacteria are essential for science and medicine,” Purcarea concluded, “but careful handling and safety measures in the lab are essential to mitigate the risk of uncontrolled spread.”
Reference: “First genome sequence and functional profiling of Psychrobacter SC65A.3 preserved in 5,000-year-old cave ice: insights into ancient resistome, antimicrobial potential, and enzymatic activities” by Victoria Ioana Paun, Corina Itcus, Paris Lavin, Mariana Carmen Chifiriuc and Cristina Purcarea, 18 December 2025, Frontiers in Microbiology.
DOI: 10.3389/fmicb.2025.1713017
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.