Scientists find variants by sequencing samples from people that have tested positive for the virus.
Since early in the COVID pandemic, the Network for Genomics Surveillance in South Africa has been monitoring changes in SARS-CoV-2. This was a valuable tool to understand better how the virus spread. In late 2020, the network detected a new virus lineage, 501Y.V2, which later became known as the beta variant. Now a new SARS-CoV-2 variant has been identified – B.1.1.529. The World Health Organisation has declared it a variant of concern, and assigned it the name Omicron. To help us understand more, The Conversation Africa’s Ozayr Patel asked scientists to share what they know.
What’s the science behind the search?
Hunting for variants requires a concerted effort. South Africa and the UK were the first big countries to implement nationwide genomic surveillance efforts for SARS-CoV-2 as early as April 2020.
Variant hunting, as exciting as that sounds, is performed through whole genome sequencing of samples that have tested positive for the virus. This process involves checking every sequence obtained for differences compared to what we know is circulating in South Africa and the world. When we see multiple differences, this immediately raises a red flag and we investigate further to confirm what we’ve noticed.
Fortunately, South Africa is well set up for this. This is thanks to a central repository of public sector laboratory results at the National Health Laboratory Service, (NGS-SA), good linkages to private laboratories, the Provincial Health Data Centre of the Western Cape Province, and state-of-the-art modeling expertise.
In addition, South Africa has several laboratories that can grow and study the actual virus and discover how far antibodies, formed in response to vaccination or previous infection, are able to neutralize the new virus. This data will allow us to characterize the new virus.
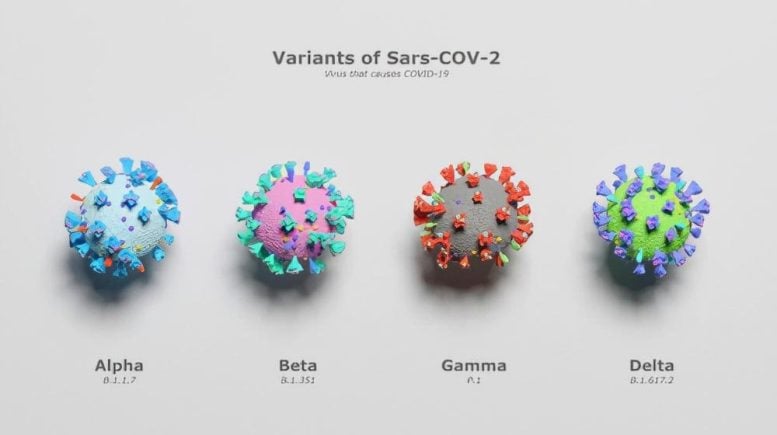
The beta variant spread much more efficiently between people compared to the “wild type” or “ancestral” SARS-CoV-2 and caused South Africa’s second pandemic wave. It was therefore classified as a variant of concern. During 2021, yet another variant of concern called delta spread over much of the world, including South Africa, where it caused a third pandemic wave.
Very recently, routine sequencing by Network for Genomics Surveillance member laboratories detected a new virus lineage, called B.1.1.529, in South Africa. Seventy-seven samples collected in mid-November 2021 in Gauteng province had this virus. It has also been reported in small numbers from neighboring Botswana and Hong Kong. The Hong Kong case is reportedly a traveler from South Africa.
The World Health Organisation has given B.1.1.529 the name Omicron and classified it as a variant of concern, like beta and delta.
Why is South Africa presenting variants of concern?
We do not know for sure. It certainly seems to be more than just the result of concerted efforts to monitor the circulating virus. One theory is that people with highly compromised immune systems, and who experience prolonged active infection because they cannot clear the virus, may be the source of new viral variants.
The assumption is that some degree of “immune pressure” (which means an immune response which is not strong enough to eliminate the virus yet exerts some degree of selective pressure which “forces” the virus to evolve) creates the conditions for new variants to emerge.
Despite an advanced antiretroviral treatment program for people living with HIV, numerous individuals in South Africa have advanced HIV disease and are not on effective treatment. Several clinical cases have been investigated that support this hypothesis, but much remains to be learned.
Why is this variant worrying?
The short answer is, we don’t know. The long answer is, B.1.1.529 carries certain mutations that are concerning. They have not been observed in this combination before, and the spike protein alone has over 30 mutations. This is important, because the spike protein is what makes up most of the vaccines.
We can also say that B.1.1.529 has a genetic profile very different from other circulating variants of interest and concern. It does not seem to be a “daughter of delta” or “grandson of beta” but rather represents a new lineage of SARS-CoV-2.
Some of its genetic changes are known from other variants and we know they can affect transmissibility or allow immune evasion, but many are new and have not been studied as yet. While we can make some predictions, we are still studying how far the mutations will influence its behavior.
We want to know about transmissibility, disease severity, and ability of the virus to “escape” the immune response in vaccinated or recovered people. We are studying this in two ways.
Firstly, careful epidemiological studies seek to find out whether the new lineage shows changes in transmissibility, ability to infect vaccinated or previously infected individuals, and so on.
At the same time, laboratory studies examine the properties of the virus. Its viral growth characteristics are compared with those of other virus variants and it is determined how well the virus can be neutralized by antibodies found in the blood of vaccinated or recovered individuals.
In the end, the full significance of the genetic changes observed in B.1.1.529 will become apparent when the results from all these different types of studies are considered. It is a complex, demanding, and expensive undertaking, which will carry on for months, but indispensable to understand the virus better and devise the best strategies to combat it.
Do early indications point to this variant causing different symptoms or more severe disease?
There is no evidence for any clinical differences yet. What is known is that cases of B.1.1.529 infection have increased rapidly in Gauteng, where the country’s fourth pandemic wave seems to be commencing. This suggests easy transmissibility, albeit on a background of much relaxed non-pharmaceutical interventions and low number of cases. So we cannot really tell yet whether B.1.1.529 is transmitted more efficiently than the previously prevailing variant of concern, delta.
COVID-19 is more likely to manifest as severe, often life-threatening disease in the elderly and chronically ill individuals. But the population groups often most exposed first to a new virus are younger, mobile, and usually healthy people. If B.1.1.529 spreads further, it will take a while before its effects, in terms of disease severity, can be assessed.
Fortunately, it seems that all diagnostic tests that have been checked so far are able to identify the new virus.
Even better, it appears that some widely used commercial assays show a specific pattern: two of the three target genome sequences are positive but the third one is not. It’s like the new variant consistently ticks two out of three boxes in the existing test. This may serve as a marker for B.1.1.529, meaning we can quickly estimate the proportion of positive cases due to B.1.1.529 infection per day and per area. This is very useful for monitoring the virus’s spread almost in real time.
Are current vaccines likely to protect against the new variant?
Again, we do not know. The known cases include individuals who had been vaccinated. However, we have learned that the immune protection provided by vaccination wanes over time and does not protect as much against infection but rather against severe disease and death. One of the epidemiological analyses that have commenced is looking at how many vaccinated people become infected with B.1.1.529.
The possibility that B.1.1.529 may evade the immune response is disconcerting. The hopeful expectation is that the high seroprevalence rates, people who’ve been infected already, found by several studies would provide a degree of “natural immunity” for at least a period of time.
Ultimately, everything known about B.1.1.529 so far highlights that universal vaccination is still our best bet against severe COVID-19 and, together with non-pharmaceutical interventions, will go a long way towards helping the healthcare system cope during the coming wave.
Written by:
- Prof. Wolfgang Preiser, Head: Division of Medical Virology, Stellenbosch University
- Cathrine Scheepers, Senior Medical Scientist, University of the Witwatersrand
- Jinal Bhiman, Principal Medical Scientist at National Institute for Communicable Diseases (NICD), National Institute for Communicable Diseases
- Marietjie Venter, Head: Zoonotic, Arbo and Respiratory Virus Programme, Professor, Department Medical Virology, University of Pretoria
- Tulio de Oliveira, Director: KRISP – KwaZulu-Natal Research and Innovation Sequencing Platform, University of KwaZulu-Natal
This article was first published in The Conversation.![]()







Be the first to comment on "The Hunt for COVID Variants: How the New Omicron Variant Was Found and What We Know So Far"