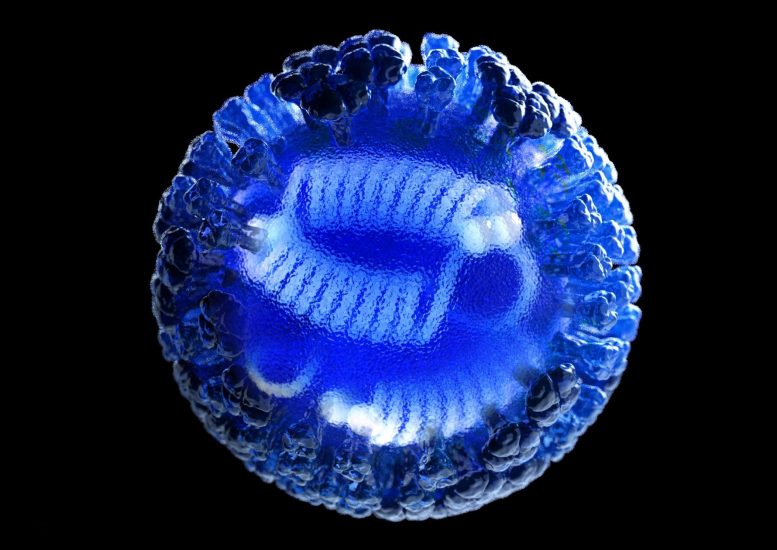
3D computer-generated rendering of a whole influenza (flu) virus. Our immune response is triggered by the virus’ hemagglutinin (HA) and neuraminidase (NA) surface proteins, shown in semi-transparent blue. HA is a trimer comprised of three subunits, while NA is a tetramer, comprised of four subunits, with a head region resembling a 4-leaf clover. Credit: Illustration by Dan Higgins, courtesy of CDC/ Douglas Jordan
The first strain of influenza virus we encounter during childhood sets the course of how our immune system responds to exposures later in life.
How successfully a person can fend off the flu depends not only on the virus’ notorious ability to change with the season, but also on the strain first encountered during childhood, according to new research published in the open-access journal PLoS Pathogens on December 19, 2019.
The findings offer an explanation for why some patients fare much worse than others when infected with the same strain of the flu virus. The results also could help inform strategies aimed at curbing the impact from the seasonal flu.
“The last two flu seasons have been more severe than expected,” says study co-author Michael Worobey, head of the Department of Ecology and Evolutionary Biology and a member of the BIO5 Institute at the University of Arizona. “In the 2017-18 season, 80,000 people died in the U.S., more than in the swine flu pandemic of 2009. Influenza is a major, major killer – not just in this country, but worldwide.”
For decades, scientists and healthcare professionals were vexed by the fact that the same strain of the flu virus affects people to various degrees of severity. Then, in 2016, a team including Worobey and authors of the current study presented a paper in the journal Science showing that past exposure to the flu virus determines an individual’s response to subsequent infections, a phenomenon called immunological imprinting.
The discovery helped overturn the prior commonly held belief that previous exposure to a flu virus conferred little or no immunological protection against strains that can jump from animals into humans, such as those causing the so-called swine flu or bird flu. These strains, which have already caused hundreds of spillover cases of severe illness or death in humans, are of global concern because they could gain mutations that allow them not only to readily jump from animal populations to humans, but also spread rapidly from person to person.
In the current study, the researchers set out to investigate whether immunological imprinting could explain people’s response to flu strains already circulating in the human population and to what extent it could account for observed discrepancies in how severely the seasonal flu affects different age groups.
The team analyzed health records that the Arizona Department of Health Services routinely obtains from hospitals and private physicians to track flu cases to study how different strains of the flu virus affect people at different ages.
Two subtypes of influenza virus, H3N2 and H1N1, have been responsible for seasonal outbreaks of the flu over the last several decades. H3N2 causes the majority of severe, clinically attended cases in high-risk elderly cohorts and the majority of overall deaths. H1N1 causes fewer deaths overall and skews more toward young and middle-aged adults.
The health record data revealed a pattern: People first exposed to H1N1 during childhood were less likely to end up hospitalized if they encountered H1N1 again later in life than people who were first exposed to H3N2. Conversely, those first exposed to H3N2 enjoyed extra protection against H3N2 later in life.
To understand the discrepancy, the researchers dug into the evolutionary relationships between influenza virus strains. H1N1 and H3N2, it turned out, belong to two separate branches, or groups, on the influenza “family tree.” While infection with one does result in the immune system being better prepared to fight a future infection from the other, the protection against future infections is much stronger when exposed to strains from the same group it has battled before.
“In other words, if you were a child and had your first bout of flu in 1955, when the H1N1 but not H3N2 virus was circulating, an infection with H3N2 was much more likely to land you in the hospital than an infection with H1N1 last year, when both strains were circulating,” Worobey says.
But the records also revealed another pattern, one that was much more difficult to explain: People whose first childhood exposure was to H2N2, a close cousin of H1N1, did not have a protective advantage when they later encountered H1N1. This seemed strange, as the two subtypes are in the same group, and the researchers’ earlier work showed that exposure to one can, in some cases, grant considerable protection against the other.
“Our immune system often struggles to recognize and defend against closely related strains of seasonal flu, even though these are essentially the genetic sisters and brothers of strains that circulated just a few years ago,” says lead author Katelyn Gostic, who conducted this research as a doctoral student in the lab of the paper’s senior author, James Lloyd-Smith, at the University of California, Los Angeles. “This is perplexing because our research on bird flu shows that deep in our immune memory, we have some ability to recognize and defend against the distantly related, genetic third cousins of the strains we saw as children.”
“Clearly, something compromises the immunity to strains that you see secondarily, even if they belong to the same group as your first exposure,” Worobey adds. “The second subtype you’re exposed to is not able to create an immune response that is as protective and durable as the first.”
In other words, our ability to fight off the flu virus is determined not only by the subtypes we have encountered over the course of our lives, but also by the sequence in which we have encountered them.
“Whichever subtype our immune system sees first lays down an imprint that protects us especially well against strains of the same subtype,” Worobey says, “but relatively poorly against strains from other subtypes, even though you’ve encountered those subsequently.”
The molecular causes of this effect are currently being studied, according to the researchers.
“Part of your immune system’s response to current infection is directed against the strain you first had as a kid, and that investment of fighting the last war appears to compromise your ability to form a fully effective immune response to the invader you encounter later,” Worobey says.
The researchers hope that their findings may help predict which age groups might be severely affected during future flu seasons based on the subtype circulating, which in turn may help health officials prepare an adequate response, such as doling out limited vaccines by cohort.
“These findings provide insight into the patterns we see in our flu surveillance and how they might change in the future,” says Shane Brady, Deputy State Epidemiologist at the Arizona Department of Health Services in Phoenix. “This highlights the importance of collaboration between public health practitioners and researchers.”
The study adds to earlier work by the same group that has made the concept of immunological imprinting a key part of the long-term battle against flu and one of the foundations of the National Institutes of Health’s strategic plan to develop a universal flu vaccine.
“We hope that by studying differences in immunity against bird flus, where our immune system shows a natural ability to deploy broadly effective protection, and against seasonal flus, where our immune system seems to have bigger blind spots, we can uncover clues useful to universal influenza vaccine development,” Gostic says.
“We need a vaccine that targets the deficits on an individualized level,” Worobey says. “Our work has clearly shown that the first virus we had can have a profound long-term effect. The bad side of that is that our immune system seems to be locked into fighting just one half of flu genetic diversity, and we need to find ways of breaking that.”
Reference: “Childhood immune imprinting to influenza A shapes birth year-specific risk during seasonal H1N1 and H3N2 epidemics” by Katelyn M. Gostic, Rebecca Bridge, Shane Brady, Cécile Viboud, Michael Worobey and James O. Lloyd-Smith, 19 December 2019, PLoS Pathogens.
DOI: 10.1371/journal.ppat.1008109
Co-authors on the study are Rebecca Bridge at the Arizona Department of Health Services, and Cecile Viboud of the Fogarty International Center at the National Institutes of Health. Funding was provided by the National Institutes of Health, the National Science Foundation, DARPA and the David and Lucile Packard Foundation.




I have two comments: It is heartening and enlightening to read about the search for a detailed atomic level chemical explanation for our immunological weakness, and the same for the strength of a particular virus, however I wish we had another equally large group of physicists looking at the very same information stream from an energetic perspective. Secondly, why is it I feel we love the easy sale of these cures to much? These discoveries result in nothing more than a financial win for a few highly motivated individuals. From that win a stagnant production, sale and advertising process emerges. We need solutions that don’t mimic the last 100 solutions, solutions that can be delivered in full as open source knowledge rather than a sale and a packaged item. We need a greater team spirit that rewards our top minds in a different manner, one that directs us away from money mining stagnation. Our present system is so wedded to a chemical sale we have forgotten where we were once headed. We need to move on faster to more knowledge, rather than wading in the sales and paybacks as if we were children waiting for our emotional storms to pass.