Heidelberg biochemists and structural biologists from Shanghai unravel the roles of two key regulatory factors in mRNA splicing.
Two molecular control factors play a key role in splicing, the process by which precursor messenger RNA (pre-mRNA) is cut and reassembled into mature mRNA, a critical step before protein production can occur in the cell. These largely uncharacterized factors are essential for ensuring the proper function of the splicing machinery. A research team led by Prof. Dr. Ed Hurt at the Heidelberg University Biochemistry Center, in collaboration with colleagues from Fudan University in Shanghai (China), has uncovered how these two cellular “quality control inspectors” operate.
Proteins, the fundamental building blocks of cells, carry out essential functions throughout the body. The instructions for building them are encoded in DNA. To translate this genetic information into proteins, the relevant DNA sequences must first be transcribed into messenger RNA (mRNA).
Initially, the cell produces a precursor mRNA (pre-mRNA) that includes both coding regions (exons) and non-coding regions (introns). Before the mRNA can be used to make proteins, the introns must be removed and the exons precisely joined together, a process called splicing, which takes place in the cell nucleus. The result is a mature mRNA strand made up solely of protein-coding exons, ready to guide protein synthesis.
The Role of the Spliceosome
Splicing is catalyzed by a large molecular machine. This spliceosome is made up of variety of protein and RNA components, and for each splicing process, it is reassembled at the sites where an exon joins an intron and the intron in turn, connects to another exon. It is absolutely vital that the splicing complex accurately recognizes exon-intron-exon junctions so the needed cuts can be made accurately.
“The precise functioning of this molecular machine has already been well researched. What remained unclear, however, was whether the spliceosome can recognize and reject a precursor mRNA with a non-authentic splice site,” explains Prof. Hurt.
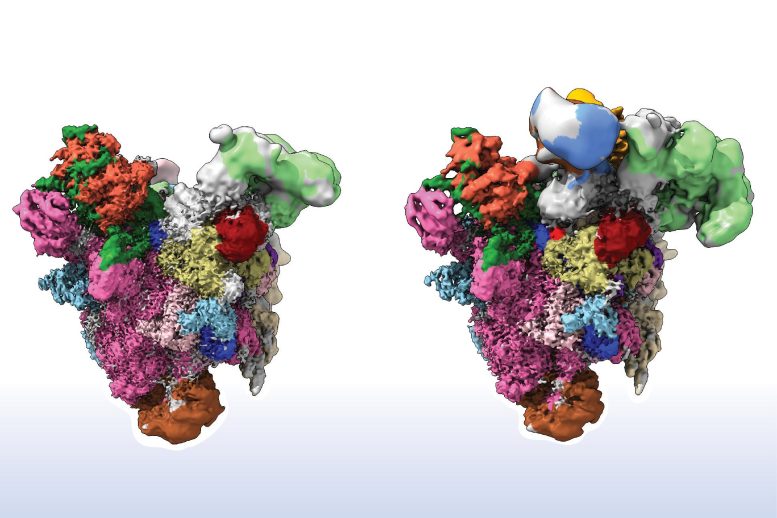
In studies with spliceosomes of the filamentous fungus Chaetomium thermophilum, the researchers were able to show that two proteins, GPATCH1 and DHX35, are crucial contributors to the precision of the splicing process. They succeeded once they were able to isolate the splicing complexes of the fungus that were in the midst of quality control and busy with rejecting a defective pre-mRNA.
Implications for Disease and Further Research
“When problems arise before the initial cut, the two proteins rush to the spliceosome to aid as quality controllers,” explains postdoc Dr Paulina Fisher. If the pre-mRNA is defective, GPATCH1 recognizes that the spliceosome should discontinue its work. As a second factor, DHX35 strips away the unsuitable precursor mRNA and eliminates it. Afterwards, the spliceosome itself is disassembled back into its individual parts, making it available for a new round of splicing.
“As cellular quality controllers, these two molecular control factors prevent a defective protein from potentially being manufactured from incorrectly spliced mRNA,” states the scientist. The researchers hope their findings will provide a better understanding of the mechanisms that ensure the accuracy of the splicing process. “They are also of clinical relevance, because defective splicing is associated with various diseases, among them cancer as well genetic and neurodegenerative diseases,” explains Ed Hurt. Along with the structural biologists from Shanghai, the Heidelberg biochemists also collaborated with a research group at the Max Planck Institute for Multidisciplinary Sciences in Göttingen.
Reference: “Structural insights into spliceosome fidelity: DHX35–GPATCH1- mediated rejection of aberrant splicing substrates” by Yi Li, Paulina Fischer, Mengjiao Wang, Qianxing Zhou, Aixia Song, Rui Yuan, Wanyu Meng, Fei Xavier Chen, Reinhard Lührmann, Benjamin Lau, Ed Hurt and Jingdong Cheng, 28 February 2025, Cell Research.
DOI: 10.1038/s41422-025-01084-w
Prof. Hurt’s research was conducted with funding from his ERC Advanced Grant. Other funds came from the National Key R&D Program of China, the National Natural Science Foundation of China, and the Shanghai Municipal Science and Technology Commission.
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.