
A new genome mapping model uncovered 62 key trait loci in bananas, overcoming chromosomal barriers and aiding future crop improvement across complex plant genomes.
Bananas are a dietary staple for millions of people, but their cultivation faces serious threats due to limited genetic diversity and significant breeding challenges. In a major scientific breakthrough, researchers examined more than 2,700 triploid banana hybrids to uncover the genetic basis of 24 important traits related to yield, plant structure, and fruit quality. By using a high-resolution SNP dataset along with an adapted genome-wide association study (GWAS) model, the team identified 62 genomic regions associated with these traits, known as quantitative trait loci (QTLs).
Many of these QTLs would have gone undetected using conventional methods because of the large chromosomal rearrangements found in banana genomes. These findings provide a valuable genetic roadmap for improving banana varieties and offer new strategies for breeding crops with complex genomes.
Breeding bananas is notoriously difficult. Most commercial bananas are sterile triploids, which means they reproduce without seeds and have limited genetic recombination, along with long and slow growth cycles. Further complicating breeding efforts, many banana varieties contain large chromosomal rearrangements that interfere with inheritance patterns and make it difficult to identify trait-linked genes.
Although there are thousands of banana cultivars globally, production is dominated by just a few, such as the widely grown ‘Cavendish’, making the entire crop susceptible to pests and environmental changes.
While GWAS has revolutionized genetic research in many crops, it has been less effective in bananas due to these genomic barriers. As global food systems face mounting challenges, understanding and addressing the complex genetics of bananas has become more urgent. To tackle this, researchers focused on developing more effective models for identifying useful genetic traits.
A new model improves trait discovery power
Published in the journal Horticulture Research, a collaborative study led by the Centre de coopération internationale en recherche agronomique pour le développement (CIRAD) investigated the genetics of 2,723 triploid banana hybrids produced from diploid and tetraploid crosses. The researchers genotyped more than 200,000 single-nucleotide polymorphisms (SNPs) and examined their associations with 24 agronomic traits using both conventional and chromosome-specific genome-wide association study (GWAS) models.
The goal was to overcome challenges posed by chromosomal translocations, which can obscure recombination patterns and reduce the ability to detect genetic associations. The new model developed by the team significantly enhanced the discovery of quantitative trait loci (QTLs), offering a clearer view of the genetic architecture behind key traits related to yield and plant structure in bananas.
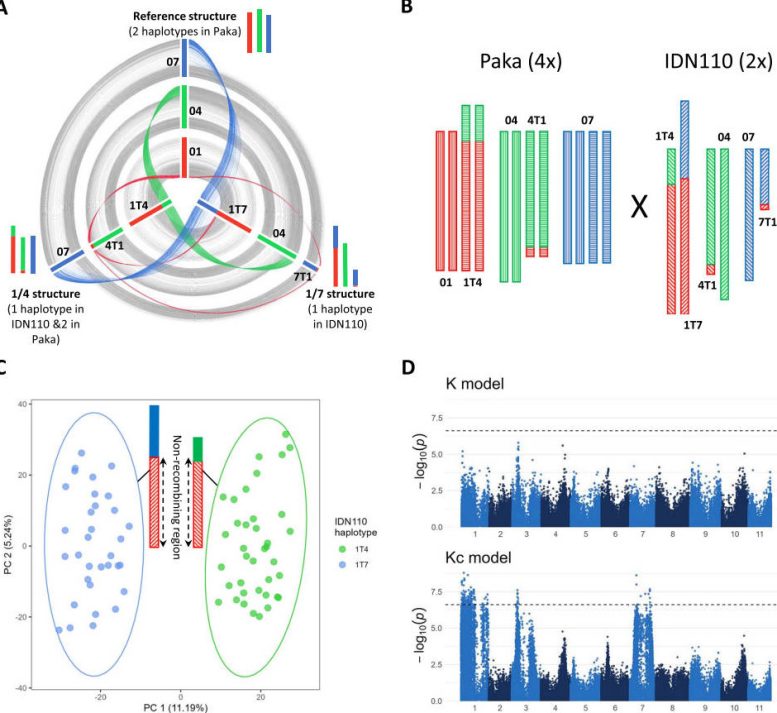
Central to the study is the “Kc model,” a novel statistical approach that improves accuracy by excluding structurally rearranged chromosomes from kinship estimation. This method allowed the researchers to detect important QTLs that the standard model had missed, including those linked to fruit size, bunch angle, and days to maturity. In total, the team identified 62 QTLs across 23 traits, many of which had not been previously mapped in bananas. Several of these QTLs showed strong ancestral connections, particularly to the banksii group, which contributed beneficial alleles for desirable traits such as shorter growth cycles and increased fruit weight.
Hotspots and multitrait selection opportunities
Meta-analysis revealed genomic hotspots controlling multiple traits, suggesting shared genetic regulation. For instance, a QTL on chromosome 3 influenced both fruit weight and bunch angle, while another on chromosome 4 affected maturity time and leaf persistence. These findings indicate pleiotropy and highlight opportunities for multitrait selection.
Notably, the study revealed how structural genome variation can both mask and mimic genetic signals, complicating trait discovery. By adjusting for these effects, the Kc model sets a precedent for future work in bananas and other crops with complex genomic architectures.
“Chromosomal rearrangements have long clouded our view of banana genetics,” said Dr. Guillaume Martin, lead author and genomics researcher at CIRAD. “This study turns that obstacle into an opportunity. By tailoring GWAS methods to banana’s unique genomic structure, we’ve uncovered important trait loci that were previously invisible. This not only benefits banana breeding—it opens a methodological path for other crops facing similar challenges. Our work shows that, with the right tools, even highly complex genomes can yield practical insights for crop improvement.”
Breeding applications and future potential
These QTL discoveries offer breeders concrete tools for improving banana varieties. Parent lines can be selected based on favorable allele profiles, especially from the banksii group. Early-genotyping of hybrids could streamline breeding by eliminating low-potential lines before field trials.
The authors also recommend pre-breeding programs focused on generating structurally homozygous lines to enhance recombination and reduce genetic drag. Beyond bananas, the study’s Kc model presents a valuable framework for trait mapping in any crop where chromosomal rearrangements limit genetic analysis. This approach could accelerate the development of high-performing, resilient cultivars to meet future food security demands.
Reference: “Genome-wide association for agro-morphological traits in a triploid banana population with large chromosome rearrangements” by Simon Rio, Lucile Toniutti, Frédéric Salmon, Catherine Hervouet, Céline Cardi, Pierre Mournet, Chantal Guiougou, Franck Marius, Claude Mina, Jean-Marie Eric Delos, Frédéric Lambert, Camille Madec, Jean-Claude Efile, Corinne Cruaud, Jean Marc Aury, Angélique D’Hont, Jean-Yves Hoarau and Guillaume Martin, 6 November 2024, Horticulture Research.
DOI: 10.1093/hr/uhae307
This research was supported by the Centre de Coopération Internationale en Recherche Agronomique pour le Développement (CIRAD) and Genoscope (from French Alternative Energies and Atomic Energy Commission (CEA)), the European Agricultural Fund for Rural Development (FEADER) and Région Guadeloupe through ‘Plan Banane Durable 1′ and ‘Plan Banane Durable 2′ programmes, the France Génomique (ANR-10-INBS-09-08) project DYNAMO, the CGIAR Research Programme on Roots, Tubers and Bananas and the Agropolis Fondation (ID 1504-006) ‘GenomeHarvest’ project through the French Investissements d’Avenir programme (Labex Agro: ANR- 10-LABX-0001-01). This work has been realized with the support of MESO@LR-Platform at the University of Montpellier and the technical support of the bioinformatics group of the UMR AGAP Institute, member of the French Institute of Bioinformatics (IFB) – South Green Bioinformatics Platform. We thank Sébastien Ricci for his thoughts on the experimental design. The authors thank Christian Vingadassalon, Frédéric Vingadassalon, Raymond Crispin, Alexin Clotaire, Ginot Karramkan, Gérard Numitor, and Nathanaelle Leclerc for their contributions to the maintenance of the experimental set-up and phenotyping. The authors thank Franc-Christophe Baurens for his biomolecular support. We thank the GPTR (Great regional technical platform) of Montpellier core facility for its technical support.
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.