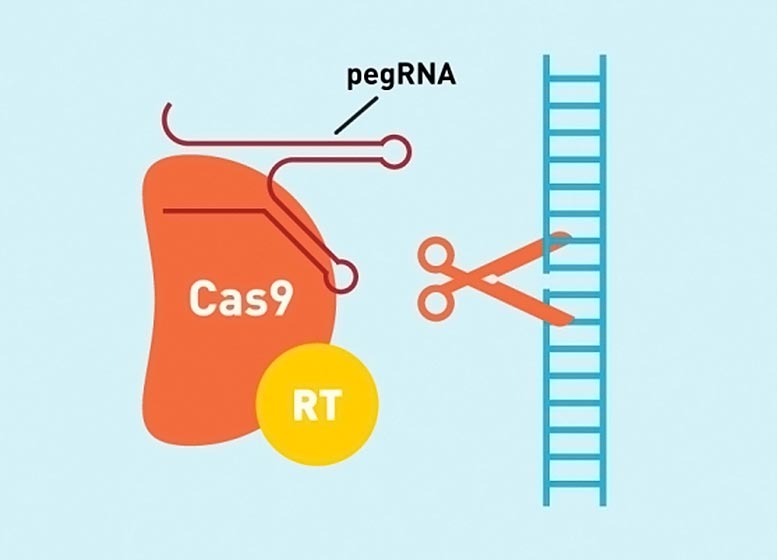
Researchers have boosted the efficiency of prime editing, a highly versatile CRISPR-based gene editing technology, and used the improved system to correct disease mutations in cells.
Scientists have developed a suite of molecular tools that increase the efficiency of a gene-editing technique called prime editing for a wide variety of cell types and target genes, expanding the scope of the technology’s therapeutic and research applications. In two new studies, the researchers used the improved prime editing systems to correct mutations linked to various neurodegenerative, metabolic, and cardiovascular diseases.
First described in 2019, prime editing is a precise gene-editing method that has the potential to correct the vast majority of known disease-causing genetic variations. Researchers can use prime editing to make DNA substitutions, insertions, and deletions at targeted sites in human cells and animals. Editing efficiency, however, varies depending on the type of cell being edited and the target location in the genome.
To further develop the technology, scientists at the Broad Institute of MIT and Harvard engineered an improvement to a key component of the prime editing system called prime editing guide RNAs, or “pegRNAs”, which encode the intended edit and direct the prime editing machinery. In a study recently published in Nature Biotechnology, the researchers showed that pegRNAs can degrade in cells, resulting in truncated pegRNAs that interfere with prime editing. They developed new pegRNAs that are protected from degradation in cells, broadly increasing editing efficiency.
In a second study published recently in Cell, Broad researchers, collaborating with scientists at Princeton University and University of California, San Francisco (UCSF), identified cellular pathways that limit prime editing efficiency, and used these insights to develop next-generation prime editing systems.
The researchers on both studies demonstrated that the new systems could more efficiently edit mutations associated with Alzheimer’s disease, heart disease, sickle cell and prion diseases, type 2 diabetes, and other diseases, while producing fewer unwanted byproducts.
“These improved prime editing efficiencies and product purities bring many edits from a regime in which they might be useful as research tools into a regime in which they may have potential as therapeutics,” said David Liu, a senior author of both studies, Richard Merkin Professor and director of the Merkin Institute of Transformative Technologies in Healthcare at the Broad Institute, professor at Harvard University, and Howard Hughes Medical Institute investigator.
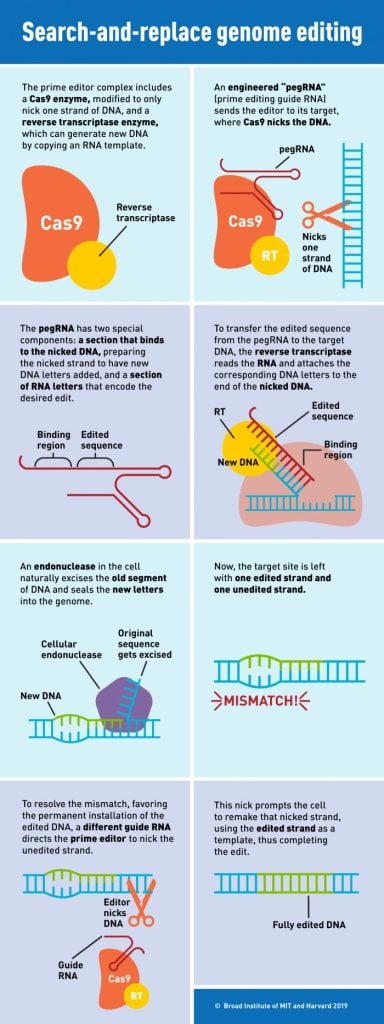
Engineering a more stable guide
Prime editing allows scientists to correct the vast majority of known disease-causing mutations — including substitutions, insertions, or deletions of up to dozens of base pairs — at specific sites in the genome. Unlike some other genome editing techniques, prime editing does not involve cutting both strands of DNA, and as a result reduces the chances of unwanted editing outcomes or undesired cellular responses. (See the infographic above for more on how prime editing works.) Hundreds of research groups are now using prime editing to study and correct mutations in a wide range of organisms including rice, wheat, zebrafish, and mice.
After first describing prime editing in 2019, Liu’s team continued to develop the technique. In the Nature Biotechnology study, they discovered a vulnerability in pegRNAs that decreased efficiency. They found that the long string of RNA at the end of the pegRNA that encodes the edit was susceptible to degradation by cellular enzymes. The degraded pegRNAs cannot mediate prime editing and also ‘poison’ the prime editing system by blocking target sites from being accessed by intact pegRNAs.
The researchers next looked for protective structures that they could add to pegRNAs. They tested several different RNA sequences, identifying sequences that folded into knot-shaped structures that shield them from RNA-degrading enzymes. When they modified pegRNAs to include the knots and a connecting sequence, they observed a substantial increase in prime editing efficiency, indicating that the new structures preserved the RNA template for editing.
Using engineered pegRNAs, or epegRNAs, in a range of mammalian cell lines, the researchers saw that epegRNAs increased prime editing efficiency three- to four-fold on average, with greater improvements in cell lines in which prime editing had previously been more difficult.
Guiding the cell towards prime edits
In the Cell study, Liu’s team and their collaborators engineered the protein component of the prime editing system to further boost efficiency and minimize byproducts produced in a broad range of cell types, including cells from patients.
The researchers aimed to understand more comprehensively the cellular factors that determine prime editing outcomes so that they could design even more efficient systems. The team suspected that certain cellular proteins active during a key part of the prime editing process — when the cell repairs DNA molecules created by prime editors — could impede or even reverse editing and increase the production of unwanted byproducts. To test this hypothesis, the researchers collaborated with teams led by Britt Adamson, an assistant professor at Princeton University; and Jonathan Weissman, a professor at UCSF when the study began and now a professor at MIT, a member of the Whitehead Institute, and a Howard Hughes Medical Institute investigator. Using CRISPR interference-based screens, the teams systematically studied the effect of turning off each of 476 different DNA repair genes on prime editing.
Based on these results, the researchers focused on a process called mismatch repair, which occurs naturally in cells to correct DNA mismatches generated during DNA replication and repair. They found that mismatch repair interferes with prime editing, decreasing editing efficiency and increasing the fraction of unintended insertions or deletions.
Armed with this insight, the team developed new prime editing systems, which they called PE4 and PE5, that include a protein, MLH1dn, that the researchers engineered to temporarily inhibit one component of mismatch repair. In cells where mismatch repair occurs, the researchers found that PE4 and PE5 substantially increased editing efficiency and produced far fewer byproducts compared to the existing prime editing systems.
Finally, the scientists created PEmax, which optimized the architecture and amino acid sequence of the prime editing machinery. Combining improvements from the PE4 and PE5 systems, PEmax, and epegRNAs resulted in a 10- to 100-fold boost in editing efficiency compared to existing systems.
“By combining the expertise of different research groups, we were able to figure out how prime editing works and optimize parts of the system,” said Adamson. “This study is a beautiful example of how fundamental understanding can drive experimental design.”
Toward therapeutics
Liu says that in many cases, the combined improvements of epegRNAs and PE4/5/max make it easier for scientists to create cell models of disease, a critical step toward developing therapeutics.
The team is now using these systems to treat cell and animal models of genetic disease, and will continue to probe the fundamental biology of these systems.
“All of these innovations are synergistic,” said Liu. “With these improvements, we’ve been able to edit important cell types with an efficiency and cleanliness that may one day help patients who suffer from diseases with a genetic component. These findings also suggest that there are other strategies out there that can further improve prime editing.”
References:
“Enhanced prime editing systems by manipulating cellular determinants of editing outcomes” by Peter J. Chen, Jeffrey A. Hussmann, Jun Yan, Friederike Knipping, Purnima Ravisankar, Pin-Fang Chen, Cidi Chen, James W. Nelson, Gregory A. Newby, Mustafa Sahin, Mark J. Osborn, Jonathan S. Weissman, Britt Adamson and David R. Liu, 14 October 2021, Cell.
DOI: 10.1016/j.cell.2021.09.018
“Engineered pegRNAs improve prime editing efficiency” by James W. Nelson, Peyton B. Randolph, Simon P. Shen, Kelcee A. Everette, Peter J. Chen, Andrew V. Anzalone, Meirui An, Gregory A. Newby, Jonathan C. Chen, Alvin Hsu and David R. Liu, 4 October 2021, Nature Biotechnology.
DOI: 10.1038/s41587-021-01039-7
This work was supported by the Merkin Institute of Transformative Technologies in Healthcare, the National Institutes of Health, the Howard Hughes Medical Institute, the Loulou Foundation, and the Bill & Melinda Gates Foundation.
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.
1 Comment
Sit back and relax, son. We’ll be at Alpha shortly.
Are we there daddy? Are we there daddy?