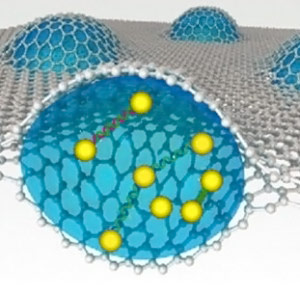
Through the use of transmission electron microscopy and a unique graphene liquid cell, Berkeley Lab researchers recorded the three-dimensional motion of DNA connected to gold nanocrystals.
Autumn is usually not such a great time for big special effects movies as the summer blockbusters have faded and those for the holiday season have not yet opened. Fall is more often the time for thoughtful films about small subjects, which makes it perfect for the unveiling of a new movie produced by researchers at the U.S. Department of Energy (DOE)’s Lawrence Berkeley National Laboratory (Berkeley Lab). Through a combination of transmission electron microscopy (TEM) and their own unique graphene liquid cell, the researchers have recorded the three-dimensional motion of DNA connected to gold nanocrystals. This is the first time TEM has been used for 3D dynamic imaging of so-called soft materials.
“Our demonstration of 3D dynamic imaging goes beyond TEM’s conventional use in seeing flat, dry samples and opens many exciting opportunities for studying the dynamics of biological macromolecular assemblies and artificial nanostructures,” says physicist Alex Zettl, one of the leaders of this research. “These results were made possible by our novel graphene liquid cell, which can meet the challenges of using TEM to image soft materials.”
Zettl, who holds joint appointments with Berkeley Lab’s Materials Sciences Division and UC Berkeley’s Physics Department where he directs the Center of Integrated Nanomechanical Systems, is one of the co-authors of a paper in NANO Letters describing this research. The paper is titled “3D Motion of DNA-Au Nanoconjugates in Graphene Liquid Cell Electron Microscopy.”
Paul Alivisatos, Berkeley Lab Director and UC Berkeley’s Samsung Distinguished Chair in Nanoscience and Nanotechnology, is the corresponding author. Other authors are Qian Chen, Jessica Smith, Jungwon Park, Kwanpyo Kim, Davy Ho and Haider Rasool.
This TEM movie (left) and 3D reconstruction (right) shows the motion of a gold nanoparticle trimer (yellow spheres) connected by DNA (green). The red arrow highlights rotational motion and dark circular shapes following this trimer are calculated projections that match what is shown in the TEM movie.
The term “soft materials” takes in a vast variety of stuff, including DNA, proteins, and other biological compounds, plastics, therapeutic drugs, flexible electronics, and certain types of photovoltaics. Despite their ubiquitous presence in our daily lives, soft materials pose many questions because the study of their dynamics at the nanoscale, especially biological systems, has been a challenge. TEM, in which a beam of electrons rather than light is used for illumination and magnification, provides the resolution for such studies but can only be used in a high vacuum as molecules in the air disrupt the electron beam. Since liquids evaporate in high vacuum, samples of soft materials, which have been described as “highly viscous fluids,” must be hermetically sealed in special solid containers (called cells) with a viewing window before being imaged with TEM.
In the past, liquid cells featured silicon-based viewing windows whose thickness limited resolution and perturbed the natural state of the soft materials. Zettl and Alivisatos and their respective research groups overcame these limitations with the development of a liquid cell based on a graphene membrane only a single atom thick. This development was done in close cooperation with researchers at the National Center for Electron Microscopy (NCEM), which is located at Berkeley Lab.
“Our graphene liquid cells pushed the spatial resolution of liquid phase TEM imaging to the atomic scale but still focused on growth trajectories of metallic nanocrystals,” says lead author Qian Chen, a postdoctoral fellow in Alivisatos’s research group. “Now we’ve adopted the technique to imaging the 3D dynamics of soft materials, starting with double-strand (dsDNA) connected to gold nanocrystals and achieved nanometer resolution.”
To create the cell, two opposing graphene sheets are bonded to one another by their van der Waals attraction. This forms a sealed nanoscale chamber and creates within the chamber a stable aqueous solution pocket approximately 100 nanometers in height and one micron in diameter. The single-atom-thick graphene membrane of the cells is essentially transparent to the TEM electron beam, minimizing the unwanted loss of imaging electrons and providing superior contrast and resolution compared to silicon-based windows. The aqueous pockets allow for up to two minutes of continuous imaging of soft material samples exposed to a 200-kilo Volt imaging electron beam. During this time, soft material samples can freely rotate.
After demonstrating that their graphene liquid cell can seal an aqueous sample solution against a TEM high vacuum, the Berkeley researchers used it to study the types of gold-dsDNA nanoconjugates that have been widely used as dynamic plasmonic probes.
“The presence of double-stranded DNA molecules incorporates the major challenges of studying the dynamics of biological samples with liquid phase TEM,” says Alivisatos. “The high-contrast gold nanocrystals facilitate tracking of our specimens.”
The Alivisatos and Zettl groups were able to observe dimers, pairs of gold nanoparticles, tethered by a single piece of dsDNA, and trimers, three gold nanoparticles, connected into a linear configuration by two single pieces of dsDNA. From a series of 2D projected TEM images captured while the samples were rotating, the researchers were to reconstruct 3D configuration and motions of the samples as they evolved over time.
“This information would be inaccessible with conventional TEM techniques,” Chen says.
This research was supported in part by the Defense Threat Reduction Agency and the National Science Foundation. The development of the liquid cell technique for in situ TEM, originally reported in the journal Science in 2012, was supported by the DOE Office of Science.
Reference: “3D Motion of DNA-Au Nanoconjugates in Graphene Liquid Cell Electron Microscopy” by Qian Chen, Jessica M. Smith, Jungwon Park, Kwanpyo Kim, Davy Ho, Haider I. Rasool, Alex Zettl and A. Paul Alivisatos, 20 August 2013, Nano Letters.
DOI: 10.1021/nl402694n
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.