UC San Diego researchers uncover enzyme behind cancer genome chaos, pointing to potential therapies for aggressive tumors.
University of California San Diego scientists have identified the enzyme that drives chromothripsis, a phenomenon in which a single chromosome breaks apart and is stitched back together in a disordered way. This extreme genetic disruption allows cancer cells to adapt quickly and develop resistance to therapy.
Although chromothripsis was first described more than ten years ago and is now recognized as a key force behind cancer progression and treatment failure, its underlying cause had remained unknown. By uncovering what initiates this process, UC San Diego researchers have resolved a major unanswered question in cancer biology and revealed new opportunities for targeting the most aggressive tumors. The study was published in Science.
Cancer cells rely on several strategies to evolve and survive treatment, but chromothripsis is notable for how much damage it can inflict at once. Rather than slowly building up genetic changes over time, this process can produce dozens or even hundreds of alterations in a single event, greatly speeding up cancer evolution. Chromothripsis is also far more common than once thought.
Scientists estimate that about one in four human cancers shows signs of this phenomenon, with even higher rates in certain diseases. Nearly all osteosarcomas, an aggressive bone cancer, exhibit chromothripsis, and elevated levels are also seen in many brain tumors.
“This discovery finally reveals the molecular ‘spark’ that ignites one of the most aggressive forms of genome rearrangement in cancer,” said senior author Don Cleveland, Ph.D., professor of cellular and molecular medicine at UC San Diego School of Medicine and member of UC San Diego Moores Cancer Center. “By finding what breaks the chromosome in the first place, we now have a new and actionable point of intervention for slowing cancer evolution.”
How Chromosomes Become Vulnerable
Chromothripsis occurs after errors in cell division cause individual chromosomes to become trapped inside tiny, fragile structures called micronuclei. Once a micronucleus bursts, its chromosome is left exposed and vulnerable to nucleases, enzymes that are capable of breaking DNA apart.
Before now, scientists didn’t know which specific nuclease triggers chromothripsis, making it impossible to target the process with cancer treatments.
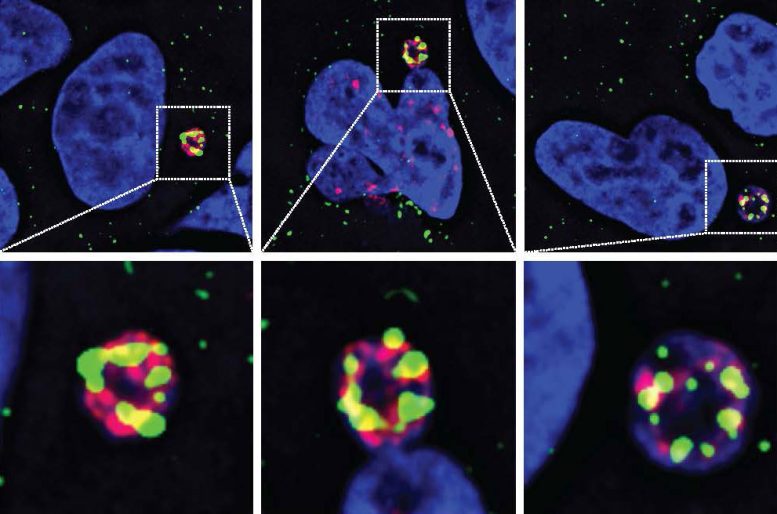
To answer this question, the researchers used an imaging-based screening technique to comb through all known and predicted human nucleases and observe how they affect human cancer cells in real time. Their analysis found one enzyme, called N4BP2, that is uniquely capable of entering micronuclei and breaking DNA apart.
To prove that N4BP2 actually causes chromothripsis, the researchers then eliminated the enzyme in brain cancer cells. They found that eliminating N4BP2 sharply reduced chromosome shattering, while forcing N4BP2 into the cell nucleus caused intact chromosomes to break, even in otherwise healthy cells.
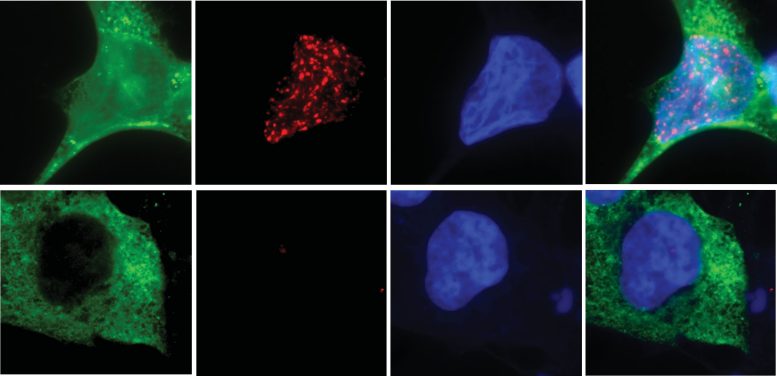
“These experiments showed us that N4BP2 isn’t just correlated with chromothripsis. It is sufficient to cause it,” said first author Ksenia Krupina, Ph.D., a postdoctoral fellow at UC San Diego. “This is the first direct molecular explanation for how catastrophic chromosome fragmentation begins.”
Links to Aggressive Tumors and Drug Resistance
The researchers also analyzed more than 10,000 human cancer genomes across many cancer types, finding that tumors with high N4BP2 expression showed significantly more chromothripsis and structural rearrangements. These cancers also exhibited elevated levels of extrachromosomal DNA (ecDNA) —circular DNA fragments that carry cancer‑promoting genes and are strongly linked to treatment resistance and aggressive growth.
Because tumors that contain ecDNA tend to be among the most difficult to treat, ecDNA has gained widespread scientific attention in recent years, including being named one of the Cancer Grand Challenges by the National Cancer Institute and Cancer Research UK. The new UC San Diego findings reveal that ecDNA is not an isolated phenomenon, but rather a downstream consequence of the much broader phenomenon of chromothripsis. By placing N4BP2 at the very start of this process, the study identifies a new molecular entry point for understanding — and potentially controlling — the most chaotic forms of genome instability in cancer.
“Understanding what triggers chromothripsis gives us a new way to think about stopping it,” said Cleveland. “By targeting N4BP2 or the pathways it activates, we may be able to limit the genomic chaos that allows tumors to adapt, recur, and become drug‑resistant.”
Reference: “Chromothripsis and ecDNA initiated by N4BP2 nuclease fragmentation of cytoplasm-exposed chromosomes” by Ksenia Krupina, Alexander Goginashvili, Michael W. Baughn, Stephen Moore, Christopher D. Steele, Amy T. Nguyen, Daniel L. Zhang, Jonas Koeppel, Prasad Trivedi, Aarti Malhotra, David Jenkins, Andrew K. Shiau, Yohei Miyake, Tomoyuki Koga, Shunichiro Miki, Frank B. Furnari, Peter J. Campbell, Ludmil B. Alexandrov and Don W. Cleveland, 11 December 2025, Science.
DOI: 10.1126/science.ado0977
The study was funded, in part, by the National Institutes of Health (grants R35GM122476, R01 ES030993-01A1, R01ES032547-01, U01CA290479-01, R01CA269919-01, R56 NS080939 and R01 CA258248).
Disclosures: Ludmil Alexandrov is cofounder, scientific advisory board member and consultant for io9; has equity; and receives income. His spouse is an employee of Biotheranostics, and he also declares U.S. provisional patent applications serial numbers 63/289,601; 63/269,033; 63/483,237; 63/366,392; 63/412,835; and 63/492,34. Andrew K. Shiau and David Jenkins are employees of FENX Therapeutics.
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.