Findings have implications for next-generation COVID-19 vaccines and treatments.
The Delta variant of SARS-CoV-2 has swept the globe, becoming the dominant variant within just a few months. A new study from Boston Children’s Hospital, published on October 26, 2021, in Science, explains why Delta is so easily spread and infects people so quickly, and suggests a more targeted strategy for developing future COVID-19 vaccines and treatments.
Last spring, study leader Bing Chen, PhD, showed how several earlier SARS-CoV-2 variants (alpha, beta, G614) became more infectious than the original virus. Each variant acquired a genetic change that stabilized the spike protein — the surface protein on which current vaccines are based. This mutation increased the variant’s ability to get into cells.
The Delta variant, which emerged soon after, is the most infectious variant known to date. Chen and colleague set out to understand why. “We thought there must be something very different happening, because Delta stands out among all the variants,” Chen says. “We found a property that we think accounts for its transmissibility and so far appears to be unique to Delta.”
Fast Fusion, Rapid Entry
For SARS-CoV-2 to infect our cells, its spikes first bind to a receptor called ACE2. The spikes then dramatically change shape, folding in on themselves. This jackknifing motion fuses the virus’s outer membrane with the membranes of our cells.
Using two kinds of cell-based assays, Chen and colleagues demonstrate that Delta’s spike protein is especially adept at membrane fusion. This enabled a simulated Delta virus to infect human cells much more quickly and efficiently than the other five SARS-CoV-2 variants (see bar chart). That was especially true when cells had relatively low amounts of the ACE2 receptor.
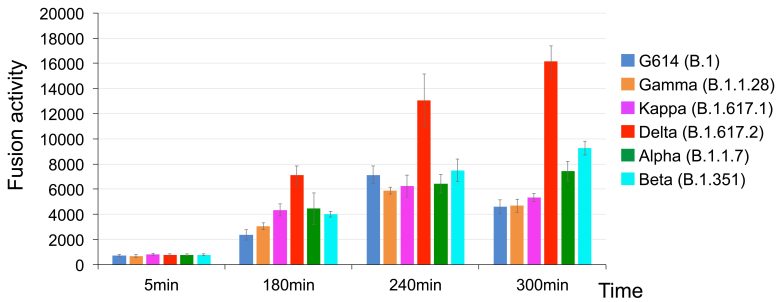
“Membrane fusion requires a lot of energy and needs a catalyst,” explains Chen. “Among the different variants, Delta stood out in its ability to catalyze membrane fusion. This explains why Delta is transmitted much faster, why you can get it after a shorter exposure, and why it can infect more cells and produce such high viral loads in the body.”
Designing Interventions, Informed by Structure
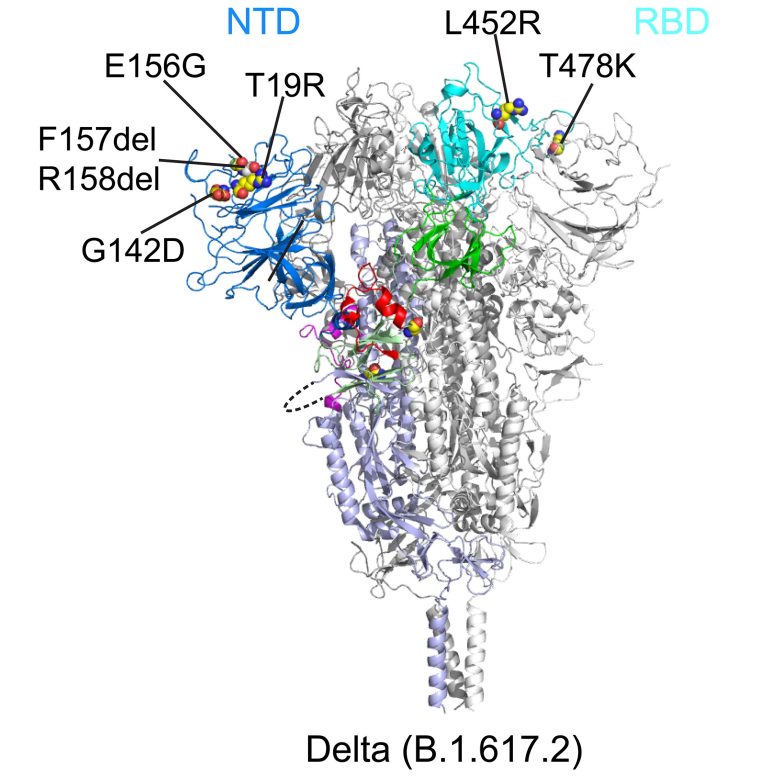
To learn how mutations in the variants affect the spike protein’s structure, Chen and colleagues used cryo-electron microscopy, which has resolution down to the atomic level. They imaged spike proteins from the Delta, Kappa, and Gamma variants, and compared them to spikes from the previously characterized G614, Alpha, and Beta variants.
All the variants had changes in two key parts of the spike protein that are recognized by our immune system’s neutralizing antibodies: the receptor-binding domain (RBD), which binds to the ACE2 receptor, and the N-terminal domain (NTD). Mutations in either domain can make neutralizing antibodies less able to bind to the spike.
“The first thing we noticed about Delta was that there was a large change in the NTD, which is responsible for its resistance to neutralizing antibodies,” Chen says. “The RBD also changed, but this led to little change in antibody resistance. Delta still remained sensitive to all the RBD-targeted antibodies that we tested.”
Looking at the other variants, the researchers found that each modified the NTD in different ways that altered its contours. The RBD was also mutated, but the changes were more limited. Overall, the RBD’s structure remained relatively stable across the variants, likely to preserve its crucial role in binding to the ACE2 receptor. The researchers therefore believe that the RBD is a more favorable target for the next generation of vaccines and antibody treatments.
“We wouldn’t want to target the NTD, because the virus can quickly mutate and change its structure; it’s a moving target,” elaborates Chen. “It might be most effective to target the RBD — to focus the immune system on that critical domain rather than the whole spike protein.”
Reference: “Membrane fusion and immune evasion by the spike protein of SARS-CoV-2 Delta variant” by Jun Zhang, Tianshu Xiao, Yongfei Cai, Christy L. Lavine, Hanqin Peng, Haisun Zhu, Krishna Anand, Pei Tong, Avneesh Gautam, Megan L. Mayer, Richard M. Walsh, Jr., Sophia Rits-Volloch, Duane R. Wesemann, Wei Yang, Michael S. Seaman, Jianming Lu and Bing Chen, 26 October 2021, Science.
DOI: 10.1126/science.abl9463
Jun Zhang, PhD, and Tianshu Xiao, PhD, of Boston Children’s Hospital were co-first authors on the paper. The study was funded by Emergent Ventures, the Massachusetts Consortium on Pathogen Readiness (MassCPR), and the National Institutes of Health (grants AI147884, AI141002, AI127193, AI39538, and AI165072).
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.
1 Comment
No surprise – the Delta variant found a better way for its spike protein to attach itself.
All the coronaviruses and all their variants have different protein spikes, with Delta having one that’s more efficient at getting around the vaccines. But the real problem is in the virus itself, not its protein shell, and why the most dangerous (MERS, SARS, and Covid-19) are so infectious. My independent research has found multiple one-in-a-million nucleotide sequence matches between all the coronaviruses and the human genome. Those sequences are the same as some of the loops of human tRNA. Using those loops and their amino acid code matches, viruses may be able to fool the nucleus membrane in cells to allow the virus to enter and associate with the human DNA, creating more opportunities for further infection. Our immune system may be compromised and may no longer be able to stop the virus and other diseases from attacking organs throughout the body. Vaccines that attack the virus protein shells while ignoring their contents are doomed to failure from the Darwin effect, but recognizing these loops suggests a possible approach to successful coronavirus vaccines. Only the infection process is considered in my work, not the innate virulence of the virus. For more info, check out this YouTube, Coronavirus – Using Your DNA Against You. https://www.youtube.com/watch?v=8dOIzD6ch8s