A new study reveals a possible protein misfolding mechanism that may resolve a long-standing mystery of why certain proteins refold into unexpected patterns.
Proteins are long molecules that must fold into precise three-dimensional shapes to function properly within cells. However, this process sometimes goes wrong, resulting in misfolded proteins that can contribute to disease if not corrected. A new study by chemists at Penn State explores a possible reason why some proteins refold into unexpected patterns.
The researchers identified a specific type of misfolding in which protein segments become improperly intertwined, creating an obstacle to normal folding. Correcting this misfolding requires substantial energy or extensive unfolding, which slows down the process and may explain the unusual folding patterns first observed in the 1990s.
“Misfolded proteins can malfunction and lead to disease,” said Ed O’Brien, professor of chemistry in the Eberly College of Science, a co-hire of the Institute for Computational and Data Sciences at Penn State, and leader of the research team. “So, understanding the mechanisms involved in the folding process can potentially help researchers prevent or develop treatments for diseases caused by misfolding.”
A research paper published on March 14 in Science Advances examines the folding kinetics of the protein phosphoglycerate kinase (PGK). The study integrates computer simulations with refolding experiments to offer a detailed analysis of the protein’s folding process.
A Different Folding Pattern: The Mystery of PGK
“For most proteins, we model the folding process as if there are two states, folded or unfolded,” said Yang Jiang, assistant research professor of chemistry in the Eberly College of Science at Penn State and the first author of the paper. “When we track the progression of a protein from unfolded to folded, we see a characteristic time-dependent pattern that we call the folding kinetics of the protein. Usually, the proportion of unfolded proteins goes down exponentially until essentially all of the proteins are folded, but some proteins don’t fit this pattern, and we were interested in the mechanisms that might explain this.”
The unusual folding pattern of PGK was first observed experimentally over 25 years ago. Whereas most proteins fit the “two-state” model of exponential folding kinetics, the molecules of PGK followed a different pattern to reach a fully folded state. This new pattern was described as “stretched-exponential refolding kinetics,” but the structural mechanism that explained this difference remained a mystery — until now. The research team hypothesized that a recently described class of misfolding may be responsible for PGK’s deviation from the traditional two-state model of folding.
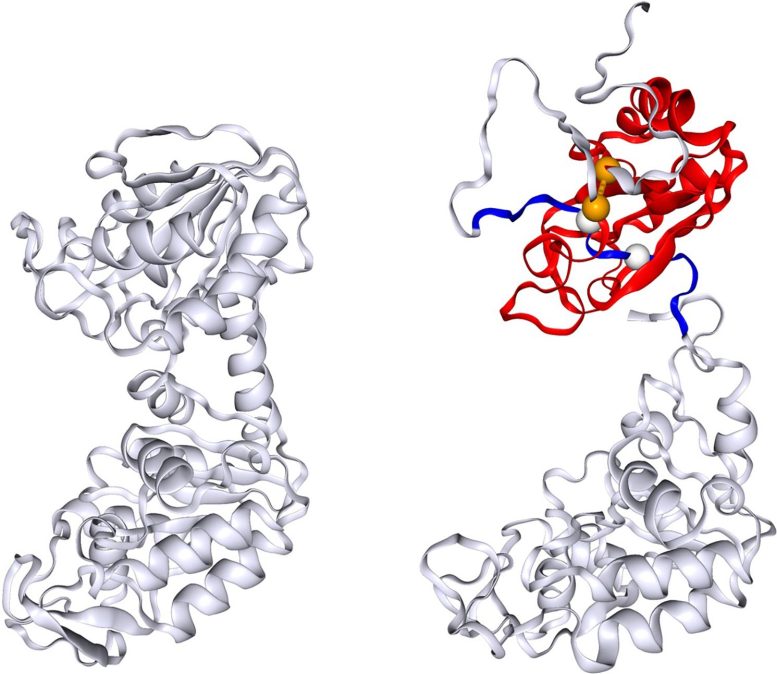
“Non-covalent lasso entanglement is a class of misfolding we recently identified where a loop of the protein traps another segment of the protein, essentially intertwining itself incorrectly,” O’Brien said. “If a protein like PGK is more prone to this type of misfolding, it could help explain why we see the stretched-exponential refolding kinetics.”
To test this hypothesis, the research team first built a computer model to simulate the folding process of PGK. Their simulations recapitulated the stretched-exponential kinetics seen in the earlier experiments. They then explored the intermediate stages of the folding process in their simulations to see if there were structural changes that could explain the stretched refolding.
Simulations and Experimental Validation
“We found several examples of misfolding involving entanglements,” Jiang said. “Sometimes a new entanglement formed and sometimes an entanglement that was part of the protein’s native structure failed to form. In our simulations, we could then remove these misfolding events and saw that the protein folded with the typical two-state exponential pattern.”
To confirm the results of their simulation, the research team, which included experimentalist Stephen Fried and lab members at Johns Hopkins University, examined the structural variation of PGK upon refolding in experiments. They found that the misfolded states predicted in the simulations were consistent with the structural signals experimentally observed in the refolded protein. They also found that these misfolded states were long-lived, suggesting that they are a crucial component of the observed stretched-exponential folding kinetics.
“Because of the nature of this type of misfolding, the protein gets stuck,” Jiang said. “The protein must backtrack in the folding process to correct the mistake, which takes time and is energetically expensive. The demonstration of this mechanism helps expand our understanding of how proteins are folded and gives an example of how it can go wrong. This is basic research, but it could eventually inform how we develop therapeutics for diseases linked to protein misfolding.”
Reference: “Protein misfolding involving entanglements provides a structural explanation for the origin of stretched-exponential refolding kinetics” by Yang Jiang, Yingzi Xia, Ian Sitarik, Piyoosh Sharma, Hyebin Song, Stephen D. Fried and Edward P. O’Brien, 14 March 2025, Science Advances.
DOI: 10.1126/sciadv.ads7379
In addition to Jiang, O’Brien and Fried, the research team includes graduate student Ian Sitarik and Assistant Professor of Statistics Hyebin Song at Penn State and co-first author Yingzi Xia and Piyoosh Sharma at Johns Hopkins University. Computer simulations and data analysis were carried using the Roar Collab, a high-performance computing cluster operated by the Institute for Computational and Data Sciences at Penn State. The U.S. National Science Foundation and the U.S. National Institutes of Health funded the research.
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.